A comparison of methods used to determine the oleic/linoleic acid ratio in cultivated peanut (Arachis hypogaea L.) ()
Peanuts are a cheap source of protein, a good source of essential vitamins and minerals, and a component of many food products. The fatty acid composition of peanuts has become increasingly important with the realization that oleic acid content significantly affects the development of rancidity. Oil content of peanuts significantly affects flavor and shelf-life. Peanut seeds normally contain between 45% - 51% oil [1]. Two fatty acids, oleic and linoleic acid, comprise over 80% of the oil content in peanut. In the fatty acid synthesis pathway, oleic acid is a precursor to linoleic acid. The enzyme responsible for the conversion of oleate to linoleate is oleoyl-PC desaturase, or Δ12 fatty acid desaturase. The fatty acid composition in most cultivated peanuts is 36% - 70% oleic and 15% - 43% linoleic acid [1]. In general, saturated fatty acids are less susceptible to oxidative degradation than their less saturated counterparts. Thus, a high oleic to linoleic (O/L) acid ratio (≥10:1) in peanut results in an increased shelf life (up to 10 times) and improved flavor when compared to a normal O/L ratio (~1.5:1). Other benefits of the high oleate trait have also been determined. Oils that have high oleic acid content and food products containing these oils have been shown to be nutritionally beneficial. Oleic acid has been shown to be associated with a reduction in blood pressure [2] and serum lipoprotein levels [3]. High-oleic peanuts have health benefits over conventional peanuts because the linoleic (polyunsaturated fat) and palmitic (saturated fat) fatty acids have been naturally replaced by the healthier oleic fatty acid (monounsaturated fat).
Since the discovery of the high O/L spontaneous mutant F435 [1], much work has been done to determine the inheritance of the high oleate trait in the different peanut market types [4-9]. Fatty acid composition of peanut is quantitatively inherited and two loci, Ol1 and Ol2, control the high O/L ratio in runner and Virginia market types [10,11]. Lopez et al. [9] confirmed that these loci also control the high oleate trait in Spanish-type peanuts. The University of Florida filed a patent on the high oleic gene in 1987 and subsequently developed and released the first high oleic cultivar, SunOleic 95R [12]. Since then rapid progress has been made towards high oleic peanut production in the US. All runner and Spanish cultivars grown in the Southwest US are high oleic. This region was first to attempt incorporating the high oleate trait into peanut cultivars, mainly to avoid off-flavors resulting from rancidity. The Southwestern US peanut industry now demands that peanuts be high oleic in nature and contracts are not offered to producers unless their product meets that requirement. The Southeast and VC production regions are making progress towards high oleic production, with high oleic cultivars accounting for a significant percentage of total production.
Selection for breeding material containing the high oleate trait is becoming easier. Traditionally, oil composition of peanut has been determined by gas chromatography (GC) analysis [13,14] which, until recently, required a fairly large sample of seed and was completely destructive, ruling out subsequent germination of tested seed. Breeders no longer have to wait for late generations to bulk up seed in order to test composite samples from advanced breeding lines for oil quality but can now begin testing as early as the F2 generation due to the adaptation of GC and development of other techniques. Oil composition of individual peanut seed can now be determined by GC analysis. Bannore et al. [15] developed an equally accurate oil extraction procedure and capillary electrophoresis (CE) protocol that enables one to extract sufficient quantities of oil from approximately 0.10 mg of peanut seed taken from the end distal to the embryo and determine the O/L ratio, ensuring subsequent seed germination and preservation of early generation breeding material. Tillman et al. [16] used near-infrared reflectance spectroscopy (NIRS) to predict oleic and linoleic acid content of single peanut seed. NIRS offers the advantage of being completely non-destructive, but lacks the accuracy that GC provides because it cannot determine exact fatty acid concentration but can only classify peanut seed as either “high oleic” or “not high oleic”. A better understanding of high oleate genetics has led to the development of several molecular marker systems which can be used to genotype F2 segregating populations and germplasm collections [4,5] and which provided a set of user-friendly high O/L molecular markers for use in marker assisted selection. Chen et al. [17] developed an allele-specific PCR assay which detects wild type and high oleic ahFAD2 alleles in the A and B genomes. Finally, Barkley et al. [18,19] recently developed a realtime PCR genotyping assay to detect the key alleles in ahFAD2 allowing for rapid high through put screening of segregating populations. This method is adaptable for seed or leaf tissue so genotyping can be performed in any generation and does not require the destruction of the seed. This method has been demonstrated to detect all possible genotypes in a segregating population [20]. The objective of this study was to analyze the accuracy of CE, two variations of NIRS, and RT-PCR, with regards to ability to determine whether an individual peanut seed is high oleic.
2. MATERIALS AND METHODS
2.1. Plant Materials
Twenty-three (23) genotypes varying in oil composition (i.e. high oleic (H) or normal/not high oleic (NH) inclusive of all four peanut market-types) were tested (Table 1). Ten to twenty seed from each genotype were chosen randomly for the method comparisons. Runner genotypes used included Tamrun 96 [21], Tamrun OL07
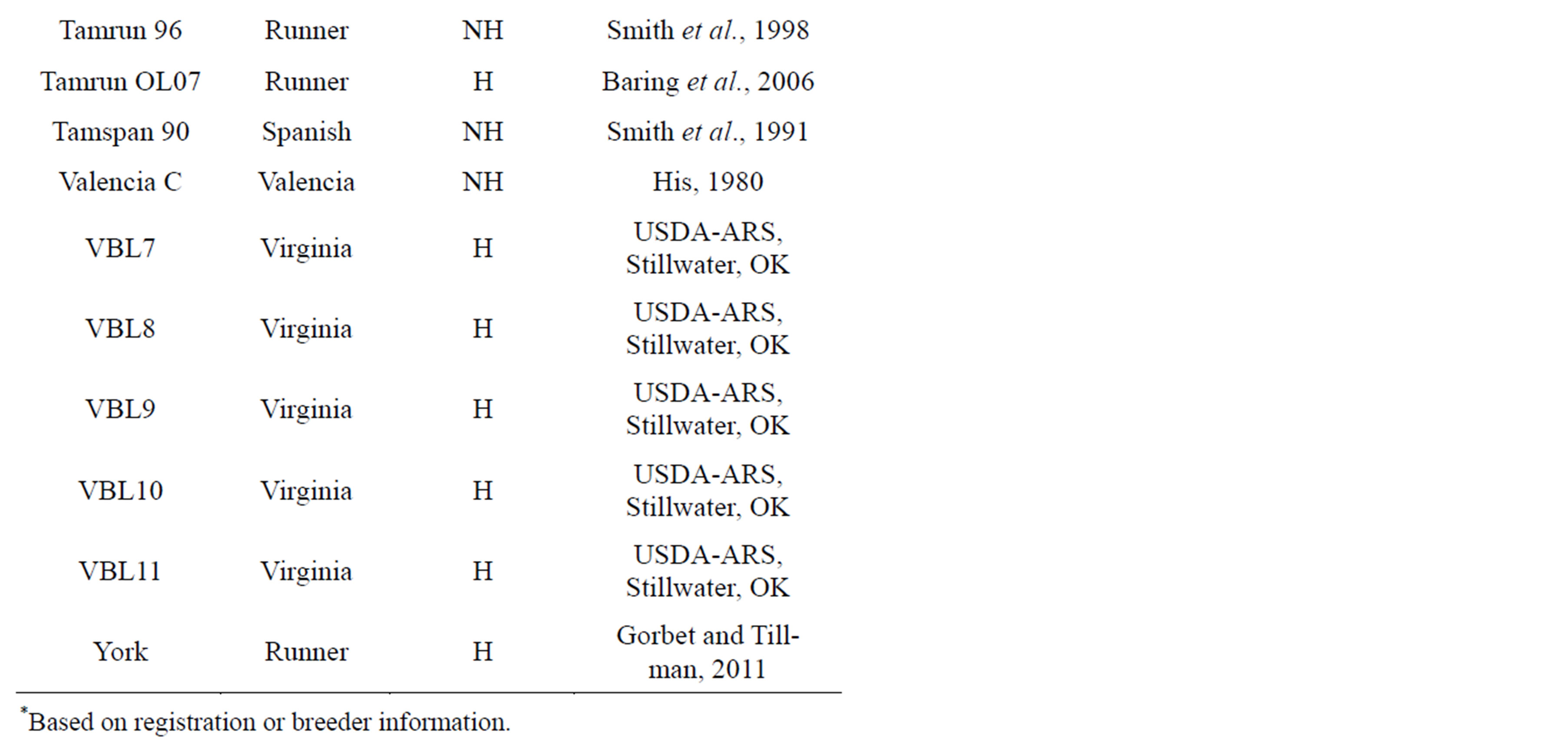


Table 1. Twenty-three peanut genotypes used in this study along with their market type, expected ranking (high oleic, H; not-high, NH) and source or reference.
[22], Red River Runner [23], Okrun [24], Georgia Green [25], AP-4 [26], Florida 07 [27], Georgia-06G [28], York [29] and McCloud [30]. Spanish genotypes used were Olin [31], ARSOK-S1 (USDA-ARS, Stillwater, OK), Tamnut OL06 [32] and Tamspan 90 [33]. Virginia genotypes tested were Brantley [11] Florida Fancy [30], Virginia breeding lines (VBLs) 7-11 (USDA-ARS, Stillwater, OK), and Jupiter (Oklahoma State University, Ag. Exp. Station, 2000). Valencia types included Valencia C [34]. No high oleic Valencia-type varieties or breeding lines were available for inclusion in this study. Samples were distributed in a blind manner, i.e. none of the researchers collecting this data knew what varieties they were testing or if they were high oleic or normal oleic. Individual seed were subjected to analysis in the following order: 1) NIRS (1 and 2), 2) RT-PCR, and 3) oil extraction. Oil extractions were then subjected first to CE then to GC.
2.2. Near Infrared Reflectance Spectroscopy (NIRS) Analysis
Two different NIRS calibration methods were applied to the samples in this study. The NIRS-1 calibration was described by Tillman et al. [16]. The NIRS-2 method was developed as follows. Two hundred and forty two seeds of various peanut cultivars and breeding lines were scanned by NIRS and subsequently analyzed for palmitic acid, oleic acid, and linoleic acid using an HP 5890A gas chromatograph (GC). The GC method described by Metcalf et al. [35] was used. Spectral data for each peanut seed were obtained from a Thermo Nicolet Industrial Solutions (Fitchburg, WI) Nexus 670 FT-IR scanning monochronometer equipped with a NearIR UpDrift Smart Accessory. Each individual seed was scanned 4 times with a mirror velocity of 1.2659 and an aperture of 10. Wavelengths were 1000 to 2500 nm. Using GC, measurements of palmitic acid, oleic acid, and linoleic acid were matched with their respective NIRS scan, and a calibration equation was developed with Thermo Nicolet Corporation TQ Analyst Professional Version 6.1.1.356 spectroscopy software. The software analysis option used to develop the equations was principal least squares (PLS). The spectra were pre-processed with mean centering, variance scaling, and a multiplicative signal correction (MSC) path length. A simple fit value algorithm (measured from zero) was used and there were 11 PLS factors for palmitic acid, 15 PLS factors for oleic acid, and 14 PLS factors for linoleic acid. The software conducted a cross-validation analysis in which each sample was left out and correlation was evaluated statistically on prediction of the left out samples. Actual versus predicted, concentration residual, and spectral residual plots were used to eliminate obvious outliers from the model. The calibration dataset was subjected to regression analysis using SAS [36] to statistically describe the relationship between the values predicted by NIRS and those obtained by GC. A separate set of 390 individual seeds from 39 various genotypes was subjected to fatty acid testing using NIRS and GC. The NIRS-predicted and GC measured palmitic acid, oleic acid, and linoleic acid values from the validation dataset were then subjected to regression analysis using PROC REG of SAS.
An important objective of screening peanut germplasm is to distinguish between “high oleic” and normal oil chemistry. Oil which is more than 74% oleic acid is considered by the industry to be “high oleic.” Therefore, observations were also made of the ability of the spectral data to correctly classify the oil when compared to the actual chemistry as determined by the GC data. In practical usage, the values predicted by the spectral data are used only as indicators of whether the oil would be classified as “high oleic” or normal. A representative graph of the NIRS-2 calibration for oleic acid is shown in Figure 1. The vertical line represents the NIRS predicted oleic value at which a peanut seed is considered to be high oleic (if greater than) or normal (if less than). The horizontal value is the GC oleic acid concentration at which a seed is consider high oleic or normal oleic.
2.3. Real-Time Polymerase Chain Reaction (RT-PCR) Analysis: DNA Extraction and PCR
All DNA samples were extracted by following the directions from an Omega-BioTek E.Z.N.A Plant DNA kit (Norcross, GA.). Slices (75 - 150 mg) from a single individual seed were used to extract DNA. Seed slices were placed in a 2 mL micro-centrifuge tube along with two 3 mm tungsten carbide beads (Qiagen Valencia, CA.) and 600 µl of P1 buffer from the Omega-BioTek kit. Tissue was pulverized by a Retsch Mixer Mill 301 (Leeds, UK) at 30 Hz for three minutes. Extracts were quantified on a DyNA Quant 200 fluorometer from Hoefer Pharmacia Biotech (San Francisco, CA). In addition, all samples were loaded on a 1% agarose gel (stained with ethidium bromide) along with a Low DNA Mass™ Ladder from Invitrogen (Carlsbad, CA) to evaluate quantity and quality of each extraction. All samples were subsequently diluted to 10 ng/µl for Real-Time PCR.
The development of the genotyping assay, the PCR master mix, and cycling conditions were all as described previously [18,19]. Genome specific SNPs identified from sequencing wild progenitors of cultivated peanut for ahFAD2 were incorporated in the probe/primer design to preferentially select the A genome when genotyping ahFAD2A or the B genome when genotyping ahFAD2B. All PCR reactions were performed in an ABI StepOne™ Real-time PCR machine using MicroAmp® fast optical 48-well plate and adhesive film seals (Applied Biosystems, Foster City, CA.). Each PCR run included non-template controls to ensure that reagents were free of contaminants. In addition, several positive controls, such as F435, were included in each run to represent the homozygous recessive mutant alleles (ol1ol2), normal oleate lines to represent the homozygous wild type dominant alleles (Ol1Ol2), and heterozygous genotypes. StepOne version 2.0 (Applied Biosystems) was utilized to analyze and score genotypes among the peanut samples and controls using the default parameters. Representative amplification plots generated from genotyping ahFAD2 in cultivated peanut using this method have been previously reported [18,19]. Alleles were detected by an increase in normalized fluorescence of Vic (green line) for the wild type alleles and an increase of fluorescence of 6-Fam (blue line) for the mutant alleles in ahFAD2A and ahFAD2B.
2.4. Fatty Acid Analysis
Saturated free fatty acid (FFAs) standards such as

Figure 1. Graph of the NIRS-2 calibration for oleic acid. The vertical line represents the NIRS predicted oleic value at which a peanut seed is considered to be high oleic (if greater than) or normal (if less than). The horizontal value is the GC oleic acid concentration at which a seed is consider high oleic or normal oleic.
stearic acid (C18:0), palmitic acid (C16:0), nonadecanoic acid (C19:0), and unsaturated FFAs such as oleic acid (C18:1), linoleic acid (C18:2), and linolenic acid (C18:3), adenosine 5’-monophosphate (AMP) monohydrate from yeast, tris (hydroxymethyl) aminomethane (Tris), acyclodextrin (a-CD) were obtained from Sigma (St. Louis, MO, USA). Dioxane, HCl, diethyl ether, and anhydrous sodium sulphate were from Fischer Scientific (Fairlawn, NJ, USA) and N-methyformamide (NMF), was from Aldrich (St. Louis, MO, USA). Hexane was obtained from EM Science (Cherry Hill, NJ, USA) and KOH was from Mallinckrodt, Inc. (Paris, KY, USA). Ethanol and methanol were purchased from AAPER Alcohol and Chemical Co. (Shelbyville, KY, USA).
2.4.1. Extraction of Oil from Individual Peanut Seed
Extraction of oil from peanut seed was performed as previously reported [15]. Seed used for these experiments were sound and mature. After removing the testa, a small portion (≤0.10 mg) of the seed was cut from the distal end (away from the embryo), weighed and ground to paste in a mortar-pestle in a sufficient volume of hexane. The slurry was transferred into a vial. The remaining paste was scraped and washed into the vial. The mortar was washed 2 - 3 times with hexane as required for complete transfer of the contents. The vial was vortexed for ca. one min and centrifuged for about 20 - 25 min at 10,000 rpm. The supernatant layer was collected and the hexane was evaporated using speed vacuum. The peanut oil left behind was further used for extraction of lipids.
The isolation of lipids from the total peanut oil was accomplished using a modification of the method described by Dermaux et al. [37]. To 20 - 25 mg of oil, 0.55 mL of 1 M potassium hydroxide in 95% ethanol solution was added and subjected to overnight reflux in a dri-bath at 97˚C. Thereafter, the mixture was cooled to room temperature, and then transferred to a separatory funnel by rinsing the reflux vial with 1.5 mL water. The non-saponifiable matter was extracted using 1.5 mL of diethyl ether. The aqueous layer containing fatty acid salts was separated from the organic layer and then acidified to pH 2.0 with 1 M HCl. The FFAs formed were extracted with 6 × 1 mL diethyl ether. All organic layers were collected, washed with 1 mL of water and dried over anhydrous sodium sulphate. The solvent was evaporated at room temperature in the fume hood overnight. The extracted FFAs were dissolved in 0.5 mL of NMF-dioxane (1:1, v/v) mixture. This was done by first dissolving the FFAs in dioxane followed by addition of NMF and then vortexing for 10 - 15 sec. The sample was divided in-half, with one portion used directly for CE analysis and the other modified for GC analysis.
2.4.2. Capillary Electrophoresis (CE) Analysis
The CE analysis of all standards and peanut samples was performed on a P/ACE MDQ (Beckman Instruments, Inc., Fullerton, CA, USA) equipped with a photodiode array detector and a 0 - 30 kV high-voltage power supply. The data were collected on an IBM personal computer configured with P/ACE MDQ 32 Karat software version 8.0. The capillary columns used for separation were untreated fused-silica capillaries with 50 mm I.D. and 363 - 359 mm O.D from Polymicro Technologies (Phoenix, AZ, USA). The total and effective lengths were 60.2 cm and 50 cm, respectively. The experiments were performed at a constant voltage of 28 KV and the temperature was maintained at 20˚C. All standards and samples were injected hydrodynamically for 3 sec by application of a pressure of 0.5 psi. The indirect UV detection was carried out at a wavelength of 254 nm with adenosine 5’-monophosphate (AMP) as a background UV absorber.
A new capillary column was flushed successively with a manual syringe using the following solutions and duration: 1 M NaOH for 10 min, followed by water for 3 min, 0.1 M HCl for 10 min, water again for 3 min, and finally with the running electrolyte for 5 min. This successive washing of the capillary was done daily at the start of the experiments using the P\ACE MDQ instrument setting and applying a pressure of 65 psi to the vial for each washing step. Before injections, equilibration of the capillary was carried out at the running voltage (i.e., 28 kV) for 20 - 30 min with freshly prepared running electrolyte solution (40 mM Tris, 2.5 mM AMP, and 7 mM of a-CD in a mixture of NMF-dioxane-water (5:3:2, v/v)).
The stock solutions of all standard FFAs were prepared by dissolving appropriate amount of acids in NMF-dioxane (4:1 v/v) to give concentration of 5 mM. All standard solutions were prepared by diluting an aliquot of the stock solutions in the running electrolyte. For the quantitative determination of oleic and linoleic acids in peanut oil samples, an internal standard that shares similar properties with these solutes is the best choice. Nonadecanoic acid (C19:0), which is usually absent or present in trace amounts in peanut oils, was selected as the internal standard for establishing the standard calibration curve. The concentrations of standard solutions for the two FFAs oleic acid (C18:1) and linoleic acid (C18:2) used for calibration were 0.2, 0.4, 0.6, 0.8, 1.0, 1.2, and 1.4 mM while the FFA C19:0 used as the internal standard had a concentration of 0.5 mM. The internal standard was first dissolved in dioxane, and then NMF was added. All stock and standard solutions and fatty acid extractions were stored at –80˚C until use. In all cases, the quantification of oleic acid and linoleic acid was achieved by comparing peak heights of the fatty acids in the sample with that of the standards from the calibration curve in the range of 0.2 mM to 1.4 mM. The calibration curves for oleic acid and linoleic acid were linear (y = 2.232x + 0.1 for oleic acid and y = 2.286x for linoleic acid) in the concentration range studied with R2 equal to 0.9985 and 0.9961, respectively.
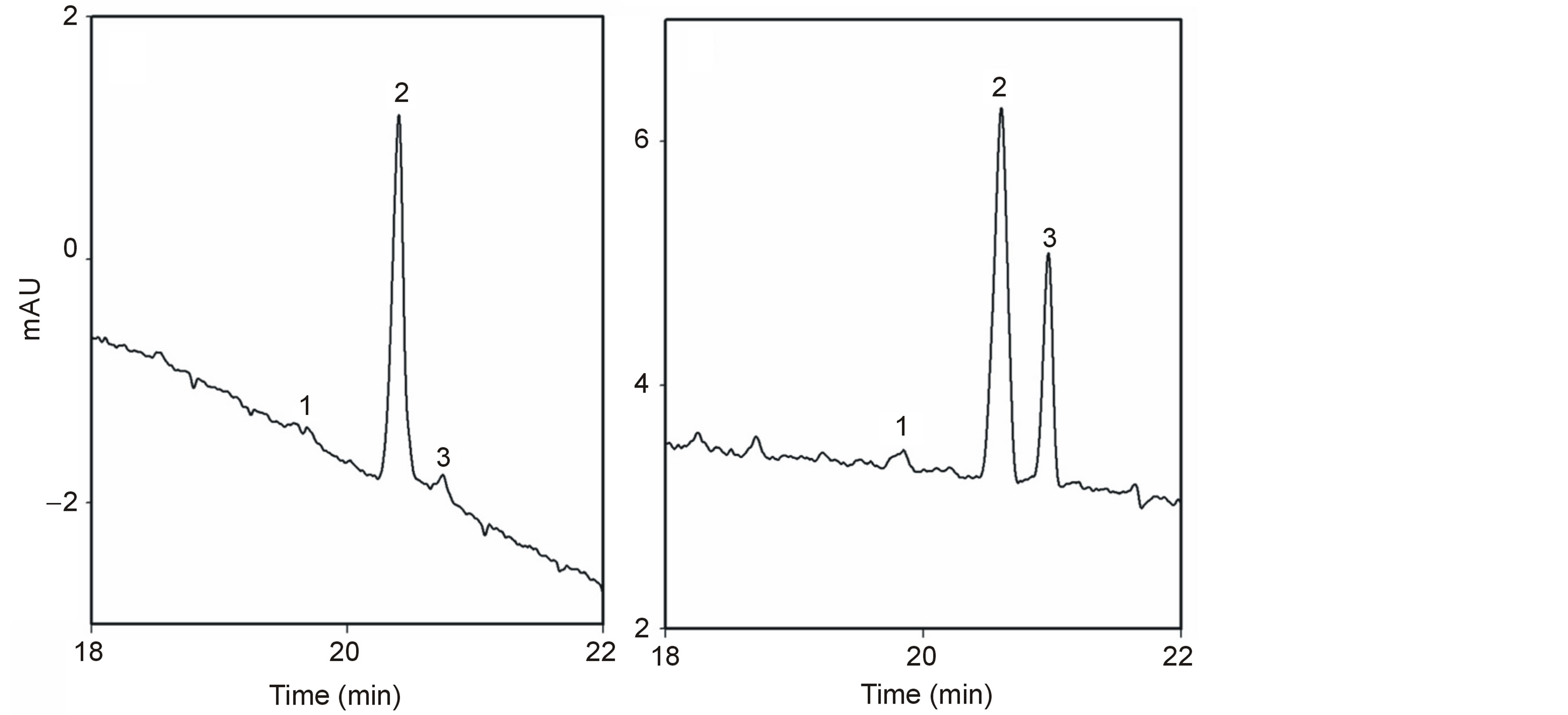
An aliquot (1 or 2 mL) of the purified peanut FFAs was taken and diluted (50-fold or 100-fold) in the final running buffer and vortexed for 4 - 5 sec. Internal standard FFA C19:0 (0.50 mM) was added to every sample. The sample was then pressure injected in the CE instrument. Hydrodynamic injection of all standards and samples was done for 3 sec at 0.5 psi (1 psi = 6895 Pa) at the anodic end. The capillary was pressure rinsed with the running electrolyte for 2 min at 65 psi between injections. Each run was performed at 28 kV with the capillary and sample temperature maintained at 20˚C, and the detection was set at 254 nm. The running electrolyte in the inlet reservoir was changed several times a day while the outlet reservoir electrolyte was changed daily. During weekends and at nights, the capillaries were stored in water. A chromatogram representative of distinguishing high oleic samples from those that are non-high oleic is shown in Figure 2.
2.4.3. Gas Chromatography (GC) Analysis
Fifty microliters of peanut extract in 1:1 n-methylformamide:dioxane was first dried under nitrogen. 50 µl of 0.5% (w/v/) butylated hydroxytoluene was added to prevent oxidation. Samples were again dried under N2 and saponified with 5% methanolic-potassium hydroxide followed by transmethylation with 14% methanolic-boron trifluoride to form fatty acid methyl esters (FAMEs) [38]. Water was removed by putting samples through magnesium sulfate mini-columns and FAMEs were purified on silicic acid mini-columns. Each sample was reconstituted in 500 µl chloroform and FAMEs were analyzed by gas chromatography (GC) on a HP5890 Series II gas chromatograph with HP7673 auto-sampler. Samples were introduced onto a DB-225 column (30 m × 0.25 mm with 0.15 µm film thickness) (J&W Scientific) using a split injector set at 250˚C with a 1:25 split ratio. Ultrapure helium was the carrier gas at 1ml/min. The GC program was as follows: 100˚C for 2 min, 25˚C/min to 180˚C, 15˚C/min to 200˚C, 4˚C/min to 225˚C and held 6 min. A flame ionization detector set at 300˚C was used and peak areas were recorded using HP-Chemstation software. FAME peaks were identified by using retention times compared to standard fatty acid methyl esters (FAMEs, NuCheck Prep, Inc., Elysian MN).
2.4.4. Statistical Analysis
The proportion of similarity of responses for each sample was calculated for each pair of methods, both across all varieties and for each variety. Using GC as the reference standard, sensitivities (correctly identifying an H sample) and specificities (correctly identifying an NH sample) were computed for the other three methods. All statistical analyses were conducted with use of PC SAS Version 9 [36].
3. RESULTS AND DISCUSSION
A total of 374 seed, inclusive of 23 genotypes and representing all four peanut market types (Table 1), were individually rated by each method as either H or NH. For
 (a) (b)
(a) (b)
Figure 2. Electropherograms from capillary electrophoresis of peanut oil samples of two lines, (a) Red River Runner (high oleic) and (b) Okrun (non-high oleic). Absorbance is measured in milli-absorbance units (mAU) at 254 nm. Peak identification: 1 = Steric acid (C18:0); 2 = Oleic acid (C18:1); 3 = Linoleic acid (C18:2).
all comparisons, the GC rating was considered “correct” since this method has been used traditionally to determine fatty acid composition in peanut oil. As expected from previous individual reports [15,16,18,35], all methods were highly successful in correctly rating individual peanut seed as either H or NH. Across genotypes, percent agreements between methods ranged from 93% - 99% with the highest being between CE and RT-PCR (Table 2). Table 3 shows a breakdown of method pair agreements in rating by genotype. All methods were in 100% agreement in their rating of seed from 9 genotypes: AP-4, Florida 07, Florida Fancy, Georgia-06G, Georgia Green, Jupiter, Okrun, Tamspan 90, and VBL11. Of those genotypes, all except AP-4 were pure (either all H or all NH) in seed composition. Average agreement of method pairs dependent upon genotype (Table 3) ranged from 93% (CE/NIRS-2 and GC/NIRS-1) to 99% (CE/RT-PCR).
Table 4 shows the agreement of method pairs and percent accuracy (compared to GC) dependent upon market type. Agreement of method pairs across market
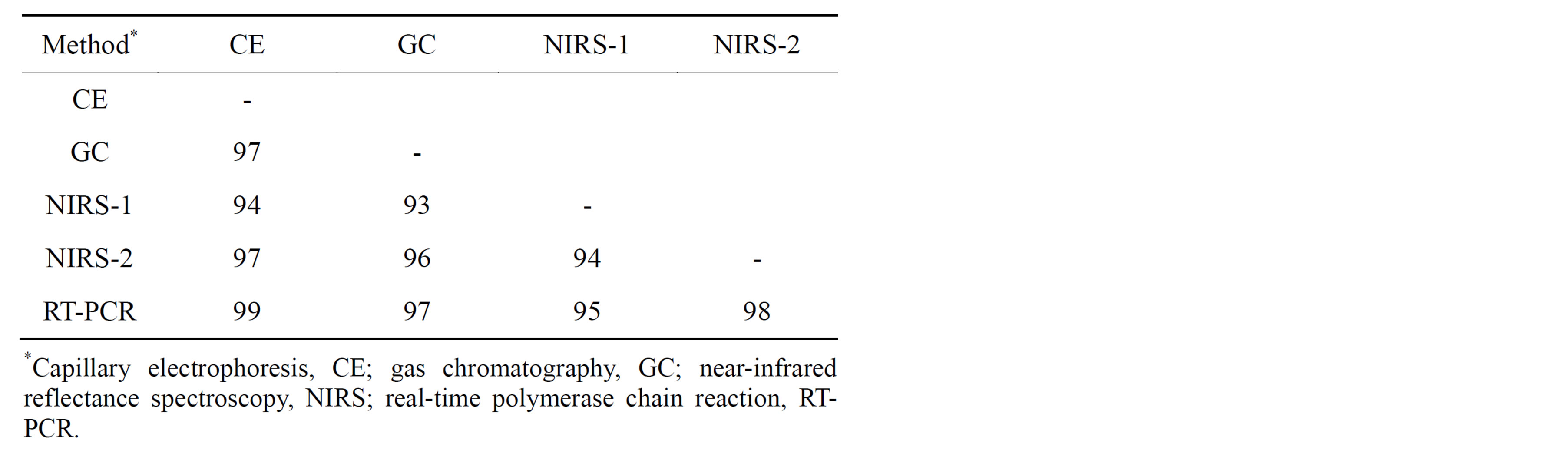
Table 2. Percent agreement of ranking high oleic (H) or not high oleic (NH) between method pairs over all peanut genotypes.

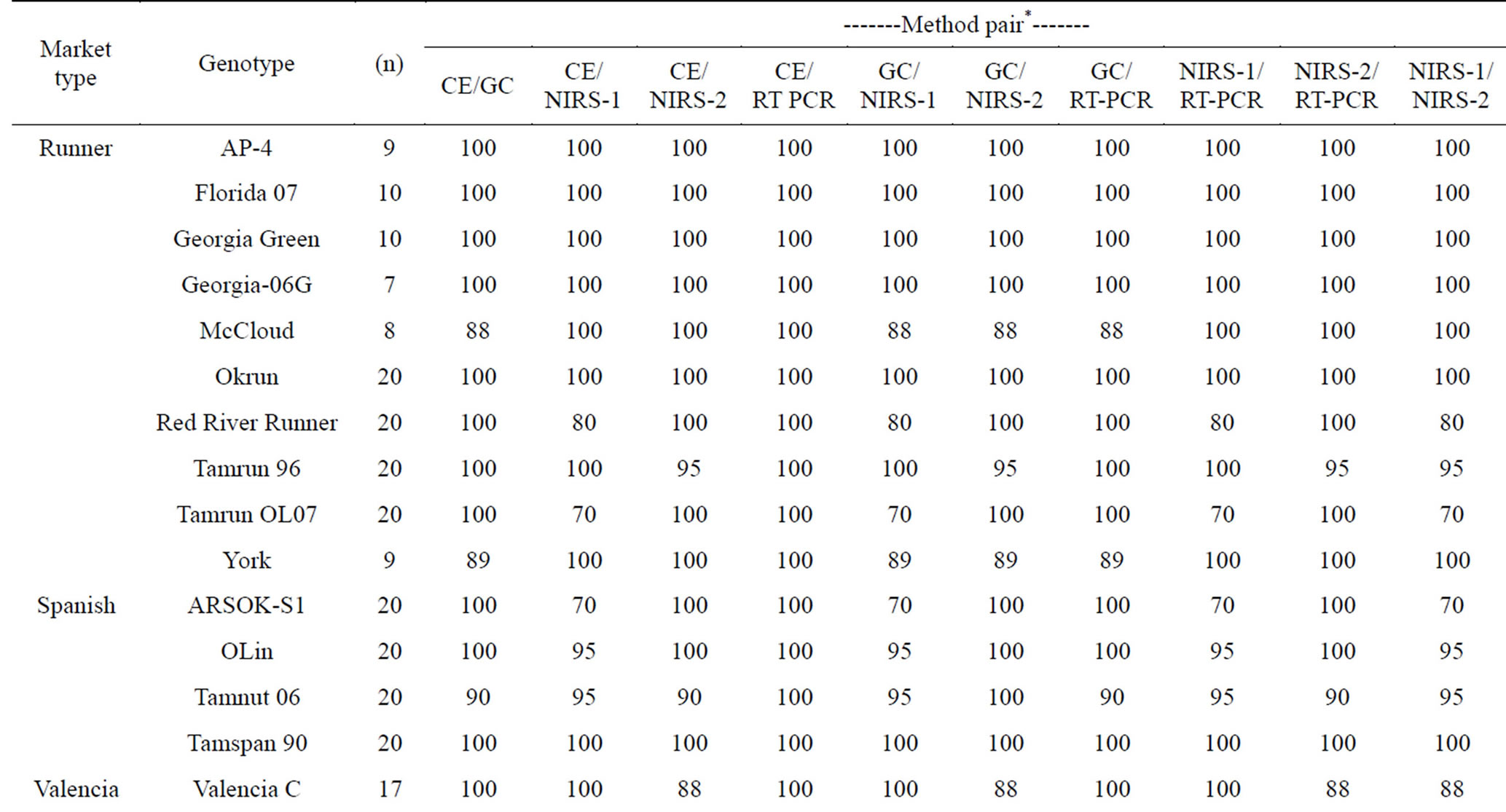
Table 3. Percent agreement of ranking high oleic (H) or not high oleic (NH) between pairs of methods dependent upon peanut genotype.

Table 4. Percent agreement of ranking high oleic (H) or not high oleic (NH) between pairs of methods dependent upon peanut market type.
type ranged from 88% - 100%, with the CE/RT-PCR method pair having an average agreement of 99%. NIRS-1/NIRS-2 was in least agreement, averaging 92% across market type. The methods most in agreement with the standard GC rating across market type were CE and RT-PCR, both averaging 98% accuracy.
Percent accuracy of individual methods (compared to the GC standard) across genotypes (Table 5) shows the most accurate rating method for predicting seed as high oleic to be CE, with 99.5%, followed by RT-PCR (98.5%), NIRS-2 (97.4%), and NIRS-1 (89.4%). For NH seed, the most accurate rating method was RT-PCR (96.2%), followed by NIRS-1 (95.7%), and CE/NIRS-2 at (94.6%).
All methods used in this study are currently employed by peanut breeding programs to assess the oleic content of early generation breeding lines. With the exception of NIRS-1, all methods have acceptable levels of accuracy (97.4% - 99.5%) in rating seed to be high oleic. Most breeding programs screen thousands of F2 seed each year and thus can afford to lose 0.5% - 2.6% of their stock due to rating error (i.e. rating a seed as NH when it is really H). Because peanut breeding programs need to subsequently germinate those seed rated as H, an accurate non-destructive rating method would be most desirable. NIRS is the only method tested here that is completely non-destructive to seed (requires no tissue be removed) and this method accurately classifies 97.4% of high oleic seed as such. Others have used NIRS to predict oil content [39] of peanuts and acid value [40] of peanut oil. NIRS can be used to predict percentages of oleic and linoleic acid in a single peanut [16], but cannot accurately determine O/L ratios since no oil extraction is performed.
RT-PCR requires removal of a small portion of seed tissue from the distal end from which to extract DNA. Alternatively, seed can be germinated and DNA can be extracted from leaf tissue, making RT-PCR a non-destructive assay. However, like NIRS, no oil extraction is performed and no prediction of O/L ratio is possible. This method, developed by Barkley et al. [18,19], rapidly identifies the wild type and mutant alleles of ahFAD2A and ahFAD2B which are responsible for normal or high oleic acid content, respectively. This method is only accurate in detecting F435 [1] derived mutants and is not effective in detecting MITE insertions in the ahFAD2 genes that can also be responsible for the high oleic phenotype [41]. Others have used PCR to study the alleles involved in the control of the high oleic acid trait [4] and the frequency of the loss of function mutation in ahFAD2A in the mini-core of the US peanut germplasm collection [5]. All of the high oleic genotypes used in this study were derived from the original donor of the high oleic trait, F435. The RT-PCR method, much like NIRS, allows many samples to be rated in a short period of time and is perfect for breeding programs needing high through-put analysis. The RT-PCR method misidentified only 1.5% of high oleic seed, an allowable loss rate for most breeding programs.
The CE method is most closely aligned with the GC method due to the fact that both directly analyze oil extracted from peanut seed for exact fatty acid composition. Comparisons of rating seed by CE and GC show the two methods to be indistinguishable in accuracy and sensitivity [42]. Similar to RT-PCR, the methods of CE, and GC are not non-destructive. Both methods require removal of a small portion of seed from the distal end for oil extraction and fatty acid purification. GC also requires modification of the fatty acids before analysis. Because of the sample preparation involved, these two methods are harder to adapt for high through-put screening. However, for programs that require an accurate O/L ratio determination for F2 seed, either of these methods will suffice. CE accuracy is dependent upon quality oil extraction and fatty acid preparation and for this reason a small error rate does exist (0.5% for CE). This study does not address error rates in the traditionally accepted GC method which is also dependent upon quality oil extraction, and thus error rates estimated for CE, NIRS, and RT-PCR may be smaller than estimated in this study.
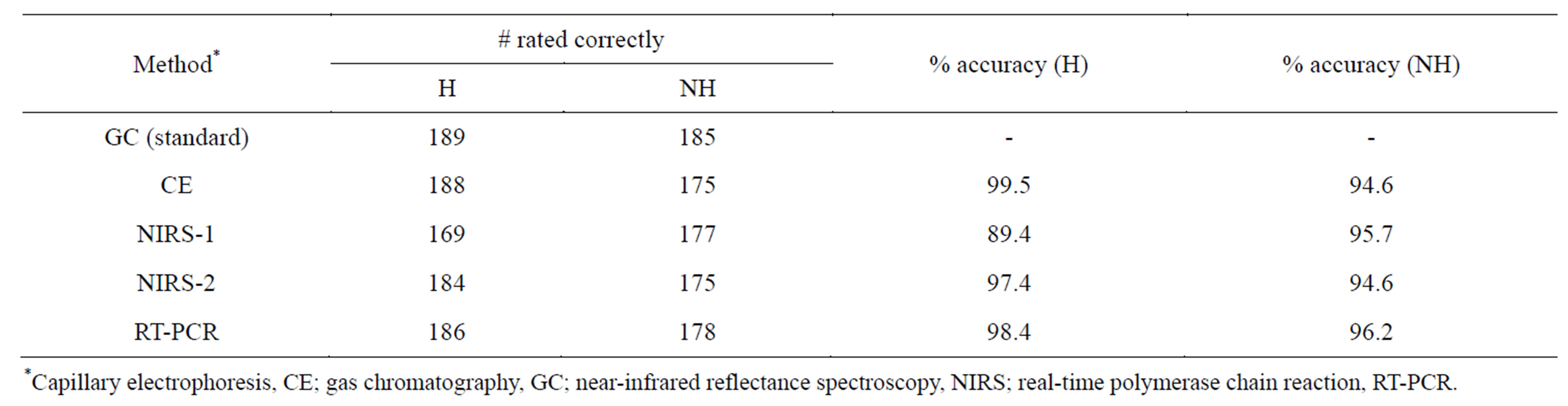
Table 5. Percent accuracy of methods compared to GC as standard reference method over all peanut genotypes.
This report evaluates the ability of techniques currently used by peanut breeding programs to accurately predict the oleic acid content of individual seed. We found that all techniques examined were extremely accurate (89.4% - 99.5%), independent of sample genotype or market type. These results indicate that peanut breeding programs can confidently use any of the methods discussed in this study for screening early generation breeding lines for oleic acid content.
ACKNOWLEDGEMENTS
The authors would like to thanks Lisa Myers and George Person for their technical assistance with this project. Funding for this project was provided by the USDA-ARS and the University of Florida. Mention of trade names or commercial products in this article is solely for the purpose of providing specific information and does not imply recommendation or endorsement by the US Department of Agriculture. USDA is an equal opportunity provider and employer.
ABBREVIATIONS
CE: capillary electrophoresis;
GC: gas chromatography;
NIRS: near infrared reflectance spectroscopy;
RT-PCR: real time polymerase chain reaction;
H: high oleic;
NH: not high oleic;
O/L: oleic acid/linoleic acid ratio.