Adaptation and Validation of a Proteolytic Activity Methodology Using Dye-Crosslinked Substrate Tablets ()
1. Introduction
In the last years, with the understanding increase of the disinfection and sterilization process in hospitals, the importance of the cleaning step in the whole process efficacy has been raised [1] [2] .
The presence of residual tissue and other body fluids on the instruments can result in layers of organic material that may be difficult to remove and disinfect, and this situation is aggravated by the formation of biofilms [3] .
Biofilms are composed by a population of microorganisms in an extra cellular matrix adhering to surfaces in which sufficient moisture is available. Bacteria growing as surface associated biofilms are much harder to treat with antimicrobials, potentially leading to dangerous infections [4] . Based on that, the importance of biofilm removal from medical and surgical instruments is known.
Disinfection processes such as heat, aldehydes or alcohols kill vegetative microorganisms but do not necessarily remove other organic contamination, including biofilm. Additionally, organic residues impair the result of low-temperature sterilization processes and heat-stable organic residues may give rise to clinical symptoms in the patient [5] .
In this context, the products employed in the organic residues removal from medical and surgical devices have gained attention, particularly the enzymatic cleaners which are responsible for organic material hydrolyses in sites of difficult access for a mechanical cleaning. These products are used to help penetration into soil on instrument surfaces, and in suspension of contaminants removed from instruments to avoid redeposition [6] . Enzymes can degrade the biofilm, by means of eroding, softening or promoting the collapse of the biofilm matrix. The biofilm removal can be achieved either by progressive erosion or by fragmented detachment [7] .
The cleaning efficacy and its products have become so important that the regulatory agency in Brazil, Anvisa, published a specific regulation for enzymatic detergents in Nov. 2012, where there are official methodologies for both proteolytic and amylolytic activities quantification [8] . These enzymatic activities need to be declared in the detergent label regarding the end of its shelf life, proving the presence of active enzymes before the expiring date. The importance of the protease was outstanding in the legislation, being required its presence in enzymatic cleaners [8] ; mainly because proteins are present in great quantities in many body fluids as blood and in biofilm as well. The official proteolytic evaluation methodology was validated by the National Institute of Health Quality Control, Oswaldo Cruz Foundation (INCQS-Fiocruz) in Rio de Janeiro, in partnership with Anvisa [8] , showing specificity, linearity, detection and quantification limits as well as precision (repeatability and intermediate precision), accuracy and robustness.
Besides the official methodology, several methods have been described for detection and determination of proteolytic activities [9] [10] . Among them, the simplest and most accessible method uses colorimetric reactions, which are detected and quantified by spectrophotometry [6] [11] [12] . The main advantage of spectroscopy is being quantitative, at a low operational cost and easy to use [13] .
Megazyme Company has several analytical kits for this purpose including Protazyme AK tablets [14] . Megazyme suggests a quantification method for protease using its tablets. In this work, some adaptation of this method using different reagents that are more available locally as well as less costly are shown.
The principle of the method is that the Protazyme AK tablets contain a blue dye-crosslinked with the protease substrate, casein, which is insoluble in the original form. With the generation of substrate fragments by the enzymatic hydrolysis, associated part of the dye becomes soluble and the solution becomes blue [14] . The advantage of this method is the visualization of the hydrolysis by the blue color intensity in the solution. Therefore, the quantification can be done by using a spectrophotometer in the visible range as well as employing a blue color intensity pattern kit. It is also possible to compare samples visually without any additional device, only by the blue color intensity, having qualitative results (Figure 1).
The proteolytic activity visualization in a simple way is very helpful during the use of enzymatic cleaners and guarantees that the product is working and also it can help in the cleaning validation in the hospital.
In this paper, a reliable method for protease activity quantification is demonstrated using Protazyme AK. This method proved to be specific, robust and repeatable. Besides that, it can be used as an alternative methodology of protease activity evaluation in hospitals, contributing with the enzymatic cleaners control and consequently the sterilization or disinfection efficacy.
2. Experimental
2.1. Materials
Protease (from Bacillus subtilis, Savinase family), amylase, cellulase and lipase were obtained from Novozymes, Protazyme AK tablets from Megazyme, hydrochloric acid p.a from Dinâmica (Brazil), tris (hydroxymethyl)- aminomethane (Tris) p.a was purchased from Nuclear (Brazil) and trichloroacetic acid p.a (TCA) from Synth (Brazil).
2.2. Apparatus
Spectrophotometer UV-Vis CECIL, CE 3021, 3000 series, using 1 cm quartz cells and absorbance (Abs) meas-

Figure 1. Qualitative visualization of the hydrolysis by the blue color intensity in the solution.
ure at 590 nm wavelength (λ); Centrifuge Excelsa II, Mod.206BL-Fanem; and Heater bath Tecnal TE-055.
2.3. Reagents Preparation
1) Tris buffer stock solution. 242 g of Tris was added to a 1 L flask with deionized water. The final volume was fixed with water after pH adjustment to 8.5 with 1 M HCl.
2) Diluted Tris buffer. 100 mL of Tris stock solution is diluted in 1 L of deionized water, with a final pH 8.3.
3) TCA reaction stop reagent. 10 g of TCA is added to a flask with 100 mL of deionized water.
4) Enzymatic standard stock solution. 15 g of protease (16 KNPU/g) is added to a flask with 100 mL of deionized water, resulting in a solution of 2.4 KNPU/ml.
5) Enzymatic standard diluted solutions. The standard stock solution is additionally diluted to 9 different concentration (0.0014, 0.0029, 0.0060, 0.0120, 0.0156, 0.0192, 0.0240, 0.0360 and 0.0480 KNPU/mL) using 100 mL flasks with 10 mL diluted Tris buffer and completing the volume with deionized water.
6) Blank solution. A solution prepared with diluted Tris buffer, with no enzymatic standard stock solution addition.
7) Enzymatic detergent solution samples. A detergent base has been prepared, containing buffer, stabilizers and surfactants, in which 15 g of protease was added (2.4 KNPU/mL), and the volume was completed to 100 mL with the detergent base.
8) Reference solution sample. A sample of a detergent base without enzyme addition.
9) Sample preparation. 1 mL of each sample solution or diluted standard solution was added to 100 mL flasks with 10 mL diluted Tris buffer, completing the volume with deionized water. The sample solution is diluted 1:100 (v:v) considering the working range to have good sensibility to read the absorbance.
2.4. Enzymatic Activity Evaluation Procedure
The procedure was based on the Megazyme assay of endo-protease using Protazyme AK tablets [14] with modifications. The Protazyme AK tablet is composed by azurine-crosslinked casein and is prepared by dyeing and crosslinking casein to produce a material which hydrates in water but is not soluble. Hydrolysis by proteases produces water soluble dyed fragments; the rate of their release is directly related to an increase in absorbance at 590 nm as well as to proteolytic activity [14] .
The main modification in the original methodology consists of changing the buffer and reaction stop reagent. The Tris buffer concentration was increased in order to reduce the pH variation with different detergent base. Besides, sodium lauryl sulphate was not added to the diluted buffer and its pH was 8.3 instead of 8.0. These changes resulted in a pH in which the used protease has a better activity. The stop reaction reagent was changed from tri-sodium phosphate solution (2% w/v, pH ~ 12.0) to trichloroacetic acid p.a (TCA, 10% w/v, acid pH), that is the stop reagent indicated by Anvisa’s methodology [8] . The other modification was the way to isolate the insoluble tablet from the reacted solution, using centrifugation instead of filtration, which was considered much more practical. Hereafter, the proposed assay procedure is described.
One Protazyme AK tablet is added to each 20 mL test-tube following addition of 1 mL of diluted Tris buffer. The tubes are heated to 40˚C, using a heating bath, for 5 minutes or until tablets disintegration. 1 mL of sample is added to each test-tube with a 30 seconds interval among each sample tube. The tubes are shaken gently. Each tube stays in the heating for 10 exact minutes, checked in the chronometer. Thereafter, 10 mL of TCA reaction stop reagent is added followed by vortex agitation. Then, samples are centrifuged at 2260 g for 10 minutes and the supernatant absorbance is read at 590 nm. The wavelength 590 nm is in the visible range and measures the intensity of blue color that can be seen in the samples. A blank was prepared using the same conditions as assay except that instead of using an enzymatic sample, it was used a buffer, as described in 2.3.6, Blank solution. The blank reading values were discounted in the read samples absorbances. The method was done in triplicate for each sample.
2.5. Validation Methodology
The method validation is based on Anvisa standard method, Resolution RE 899 of May, 29th, 2003―Guide for Analytical and Bioanalytical Methods Validation [15] .
The parameters recommended to be obtained by test type, component quantification, are: robustness, specificity, linearity, precision and intermediate precision as well as accuracy. Additionally, the detection and quantification limits were calculated. The related procedures are described, as follows.
1) Specificity. The specific action of the protease under the substrate hydrolysis was tested by the presence of other compounds in the samples, as the detergent base; other enzymes: amylase, lipase and cellulase; and impurities, denatured enzymes. The samples were tested by comparing the absorption at 590 nm of the standard sample with protease enzyme only. Means were compared by statistical analysis.
2) Robustness. The influence of many different possible variables was evaluated in the methodology: solution base type, pH, temperature and time of reaction. The absorbance means and standard deviations were calculated and compared graphically.
3) Linearity. The linearity curve range was determined by evaluating 9 points of proteolytic activity, from 0 to 0.048 KNPU/mL, being the point zero, the blank solution. The result is a mathematical expression used for the calculus of enzymatic activity in the samples. The obtained Equation (1) is represented by the expression:
 (1)
(1)
beingy: measured absorbance;
x: proteolytic activity;
a: intersection with y, when x = 0;
b: slope of the line (sensibility).
4) Limits of detection and quantification. The limits were defined from statistical calculus by average slope and activity standard deviation of 3 linear curves. Following the Equations (2) and (3):
 (2)
(2)
 (3)
(3)
beingDL: detection limit;
QL: quantification limit;
S: standard deviation of the intersection with y-axis;
b: slope of the line.
5) Precision and intermediate precision. Precision was found assaying in the same way 10 sample solutions in the same day. Intermediate precision was evaluated repeating the assay with 6 sample solutions on a different day as well as operator. The intermediate precision standard deviation was calculated by the following Equation (4) obtained from ISO 5725-3 [16] :
 (4)
(4)
whereSi: intermediate precision standard deviation;
j: the sample number, being t the number of samples;
k: the number of assays;
yjk: result of sample j and the assay k.
6) Accuracy. Samples of 3 different known proteolytic activities in a detergent base were prepared and assayed in triplicate. The activity results were compared to the known values.
2.6. Statistical Analyses
All the statistical analyses and charts were obtained by using Minitab Software, however intermediate precision was calculated by Equation (4).
3. Results and Discussion (Validation Studies)
3.1. Specificity
The specificity study results are presented, as follow.
3.1.1. Other Compounds Present in Formulation Influence
In order to verify the best wavelength to read the absorbance of the analysis, wavelength scans were performed reading samples of blanks and protease standards in buffer (Tris) and detergent base as well. Figure 2 shows the maximum absorbance peak at 590 nm, visible blue color reading, without significant interference of other detergent base components. Considering this results, 590 nm was defined as a feasible wavelength for absorbance reading, in which were obtained all the following sample results.
3.1.2. Other Enzymes Presence Influence
The protease activity alone was compared to amylase, lipase and cellulase activities in order to demonstrate that only the protease acts on the hydrolysis of azurine-crosslinked casein, resulting in blue color.
The evaluated samples were prepared by adding the other enzymes in the enzymatic standard stock solution (sample: enzymes) and by preparing a stock solution with only lipase, cellulase and amylase (sample: other enzymes).
Figure 3 shows the absorbance results of the evaluated samples. Both samples with protease have the same activity, 0.024 KNPU/mL, demonstrating that the other enzymes do not interfere in the result. This result is in agreement with our expectation considering that the enzymatic action is specific to the corresponding substrate.
Hence, the method can be considered selective in relation to the most usual enzymes found in enzymatic detergents of hospitals. The enzymes amylase, cellulase and lipase do not change the blue color intensity measured by spectroscopy, as shown in Figure 3.
3.1.3. Impurities Presence Influence
The presence of impurities was obtained by heating the enzymatic standard stock solution with addition of the enzymes amylase, lipase and cellulase to 100˚C for 5 minutes, leading to the enzymes denaturation and impurities generation. After cooling to room temperature, protease was added to 0.024 KNPU/ml in part of the sample, identified as impurities + protease; another part was measured without extra protease addition, called impurities. As a reference, enzymatic standard solution was prepared without being heated. The protease activity addition is the same in all samples.
Figure 4 shows the absorbance results of samples with or without impurities.
The most probable impurities in an enzymatic cleaner, denatured enzymes, do not affect significantly the proteolytic activity determined by the proposed methodology, as shown in Figure 4.
Thus, the studied parameters, other enzymes and impurities presence, demonstrate that the method is specific,
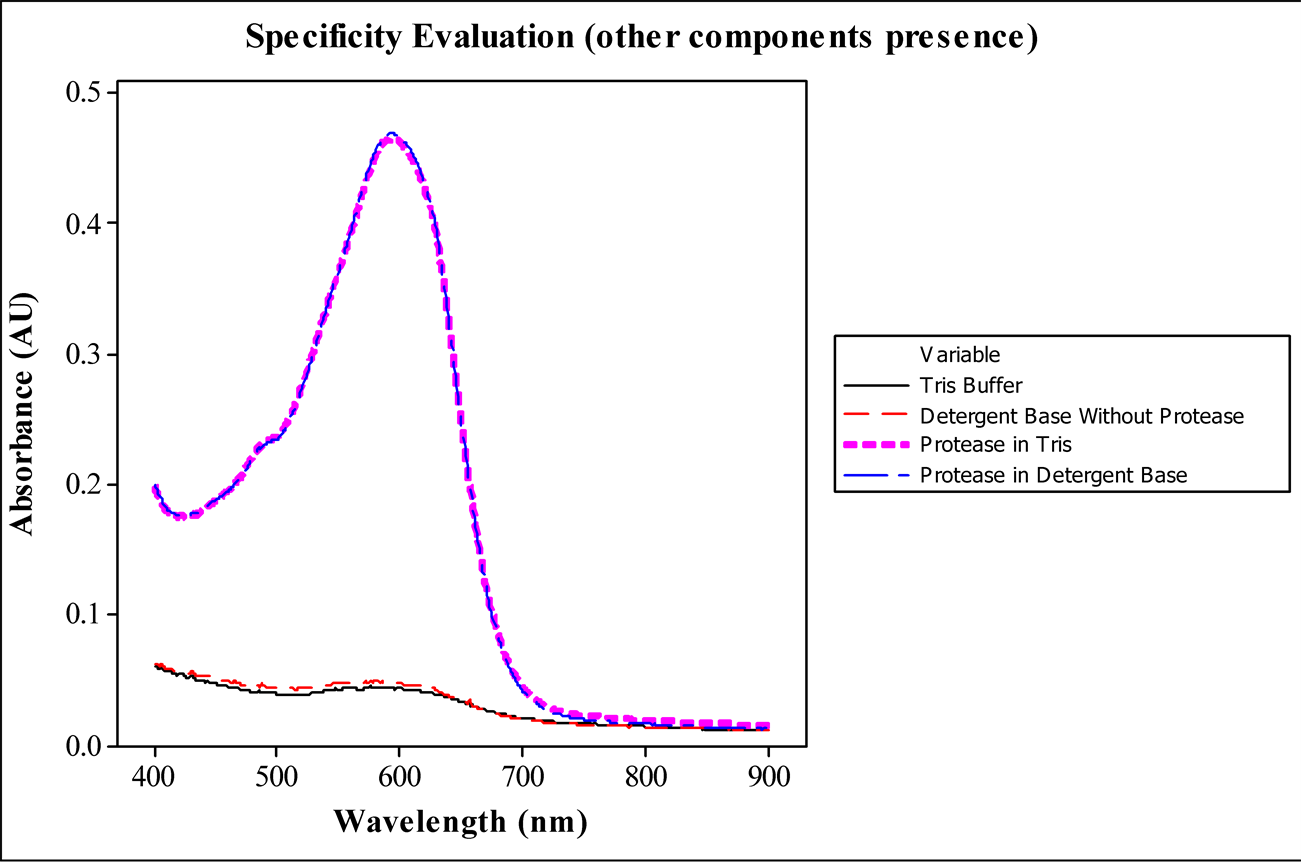
Figure 2. Evaluation of the protease specificity and determination of wavelength for sample reading by wavelength scan of analysis samples: Tris buffer, detergent base, protease in Tris and protease in detergent base.

Figure 3. Evaluation of the protease specificity by amylase, lipase and cellulose presence, being Abs0 the blank sample absorbance, and Abs the prepared sample absorbances. Enzymes: sample with protease, amylase, lipase and cellulase; other enzymes: amylase, lipase and cellulase; standard protease: enzymatic standard stock solution.
with no influence of other compounds in blue color generation.
3.2. Robustness
The parameters to be evaluated in the robustness study were chosen based on the knowledge about enzymatic activity main influences, as: pH, temperature and reaction stop time.
3.2.1. Solution Base Type and pH Influences
The influence study of solution base type and pH were done comparing samples of enzymatic detergent solution and enzymatic standard stock solution, both containing the same protease activity (2.4 KNPU/ml). The sample preparation was concluded diluting 1 mL of these solutions in 10 mL of diluted Tris buffer with different pHs (8.3, 8.4, 8.5 and 8.6) and completing the 100 mL flask with deionized water.

Figure 4. Evaluation of the methodology specificity by impurities presence, being Abs0 the blank sample absorbance, and Abs the prepared sample absorbances. Impurities: thermal denatured enzymes; impurities + protease: thermal denatured enzymes with extra protease addition; standard protease: enzymatic standard stock solution.
Instead of studying the influence of different solvent brands, recommended by RE 899 [15] , the influence of solution base was studied, because this condition was evaluated to be more critical to the methodology. The presence of a different buffer in the detergent base sample can lead to a greater pH difference than a different brand of the same buffer. The pH is a critical parameter in the enzymatic activity since the activity has a pH optimal range.
Figure 5 shows the absorbance results and deviations of the study.
The results show a significant difference between pHs 8.3 - 8.4 and 8.5 - 8.6 for the detergent base and between pHs 8.3 - 8.5 and 8.6 in the Tris buffer base. Considering the differences between the two sample basis kinds, the differences increase with the pH. For pH 8.3, it is not observed significant difference between the bases. However, the standard deviations are higher for the detergent basis. Probably, the final pH of the reaction solution is less stable by the presence of another buffer in the sample. At higher pHs, the initial diluted buffer (pH 8.3) had a greater addition of sodium hydroxide solution to achieve the pH of the test, which may also have led to higher pH ranges. Moreover, the detergent base has an alkaline buffer, which intensifies the pH increase, leading to a more pronounced increase in the absorbance intensities to the detergent base samples.
According to the pH influence study, the hydrolysis rate is increased from pH 8.3 to 8.6, what is expected considering the optimum pH activity of Savinase, over pH 9. Hence, the pH adjustment needs to be precise and the same diluted solution preparation must be employed in all the samples in the same assay.
3.2.2. Influence of the Reaction Temperature
Figure 6 shows the absorbance results of the same sample reacted with temperature variation.
Although the protease activity temperature dependency is known, in the tested range it was not statistically significant, as shown in Figure 6.
3.2.3. Reaction Stop Time Influence
The study of reaction time influence was conducted by using the same samples and preparations and stopping the reaction with TCA in different times. Figure 7 presents this study absorbance results.
In the robustness evaluation of reaction stop time, results in Figure 7, there is a significant increase in the absorbance intensity in the range from 8 to 9 minutes, in which the increase begins to become smaller until it reaches constant values at approximately 11 minutes of reaction. Possibly, this result shows that the hydrolysis of casein substrate by the protease is close to its saturation from 10.5 minutes, since it has not achieved the equipment maximum absorbance yet, which is reached over 1. It is also verified that 30 seconds of difference from 10 minutes, 9.5 and 10.5, there is not a statistical significance among the reaction times. Then, 10 minutes of reaction stop time is a feasible condition to have less influence of the time in the absorbances and cones-

Figure 5. Evaluation of the methodology robustness by diluted buffer pH variation and different base solution, being Abs0 the reference sample absorbance for detergent solution, and blank sample for standard stock solution; and Abs the prepared sample absorbance.

Figure 6. Evaluation of the methodology robustness by temperature variation, being Abs0 the blank sample absorbance, and Abs the prepared sample absorbances.
quently in enzymatic activities results.
Considering all parameters in the robustness evaluation and comparing the relative standard deviation (RSD) of the means for each studied variable, the most critical variable in the tested ranges is the base solution, being RSD over 10%, which is the most difficult to control as well. Hence, in order to increase the method accuracy, a standard curve needs to be sampled in each assay, containing the same base solution of the samples to be analyzed. This procedure may decrease the influence of base solution type in the activity determination.
3.3. Linearity
The linearity was studied in a large range, larger than 80% to 120% of the active agent, because it needs to be general for different protease activities present in different enzymatic detergents, being this range the recommended working range. Samples with higher proteolytic concentrations than the working range need to be diluted to the limits presented in this study and showed in the Figure 8 and Section 3.4 (QL value) for evaluation. The de-

Figure 7. Evaluation of the methodology robustness by reaction stop time variation, being Abs0 the blank sample absorbance and Abs the prepared sample absorbances.
fined working range leads the methodology to acceptable accuracy level as showed in Section 3.6, Table5
Figure 8 shows the measured absorbance profile in relation to the protease activity, from where it was possible to define the detection as well as the quantification limits.
Statistical calculi were done using the obtained profile to verify its linearity. The following data was determined: correlation coefficient (r), intersection with y-axis (a), slope of the line (b), sum of squared residuals of linear regression (SS); the F ratio (F) and the probability value (p) as well. The y-variables were obtained by the difference between each sample and the blank measured absorbance.
Through analysis of variance the linearity of method can be tested as well as the statistical significance of the plotted curve [17] . Table 1 shows this analysis, being DF: degree of freedom and MS: mean square error.
From the F-test table, Fcritical is 4.2, hence F calculated (Table 1) is much higher than Fcritical, being the regression hypothesis not null. Additionally, the p result shown in Table 1, lower than 0.05, indicates that the linear regression data is valid and significant with 95% of confidence. So, it was possible to obtain a standard curve for the activity. The regression parameters obtained by the minimum square method are presented in Table2
Evaluating the probability values of null hypothesis in the Table 2, it was concluded that the equation parameters a and b were valid.
Hence, the follow regression equation was determined as the standard proteolytic activity profile (Equation (5)):
 (5)
(5)
where: r = 0.995.
3.4. Limits of Detection and Quantification
The detection and quantification limits were calculated from the linear regression, by using Equations (2) and (3), respectively, being S = 0.010.
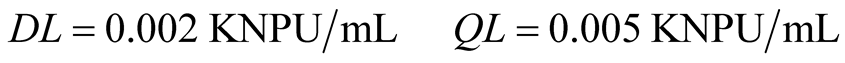
3.5. Precision and Intermediate Precision
Precision or repeatability was obtained with a fixed proteolytic activity in the average of the working range, on the same day and operator. The results are presented in Table 3, where the relative standard deviation can be seen.
Intermediate precision repeated the precision procedures on a different day and operator, its results were calculated by expression (4), considering 6 samples (t) and 2 assays, and are shown in Table4

Figure 8. Absorbance profile in relation to proteolytic activity, being Abs0 the blank sample absorbance, and Abs the prepared sample absorbances.

Table 1. Analysis of variance (ANOVA).

Table 2. Regression parameters of minimum square method.

Table 3. Results of method repeatability for proteolytic activity.

Table 4. Results of method intermediate precision.
The results shown in Table 3 and Table 4 demonstrate that the variation grade of the proteolytic activities (relative standard deviation) using the proposed method are acceptable for both the repeatability and intermediate precision, considering the RE 899/2003 (maximum 5%) [15] .
3.6. Accuracy
In order to obtain the accuracy of the method, samples with 3 different known enzymatic activities were prepared adding the protease in detergent base solution. The results were obtained calculating the recovery activity in relation to the expected values and are shown in Table5 Each activity was evaluated in triplicate.
The recovery percentage is acceptable, being in the range of 95% - 105%. Thus, the method can be considered accurate.
4. Conclusions
The proposed method was validated and the robustness, specificity, linearity, precisions and accuracy were dem

Table 5. Evaluation of the accuracy of the proposed method.
onstrated. Therefore, Protazyme tablets can be considered good substrate for protease, resulting in blue color intensity with good relation to the enzymatic activity, being adequate for analysis of enzymatic cleaners, including aging studies.
Despite the presented method being validated in specific and controlled conditions, its evaluation demonstrates that the Protazyme tablets have reproducible results and can be used for visual comparison among different samples in Central of Sterilization Supply Department of hospitals, as the absorbances are measured at a visible wavelength, although in this case there will not be quantitative results.
Thus, the method is appropriate for quantification of proteolytic activities. In addition, the used tablets can be very useful to help in enzymatic detergent evaluation in hospitals, visually comparing products in real use conditions.