Effects of Preparation and Storage of Agar Media on the Sensitivity of Bacterial Forward Scattering Patterns ()
1. Introduction
Today a large amount of our food is being processed and sold as ready-to-eat product on the shelves of the grocery stores worldwide. Foods are consumed within few days to few hours after being harvested to minimize food spoilage and contaminations [1]. To maintain food quality and safety, scientists apply various quality control strategies; however, lack of rapid, user-friendly and high through put techniques delays quality assurance in food production and processing practices. Such constraints may increase the risk of foodborne outbreaks [2,3]. Food products are highly susceptible to contaminations if they are not processed under good manufacturing practices (GMP) or following HACCP plans [4,5] . Thus, the detection of foodborne pathogens is vital and critical for the food industry to deliver and maintain food safety [6]. Recent E. coli O104-outbreak in Europe [7] in 2011 and Salmonella in the United States [8] emphasize the need for an improvement of current culture-based and direct detection techniques of analysis; and development of new technologies to mitigate foodborne outbreaks and resulting economic damages.
In this context, a novel label-free light scattering sensor can deliver rapid and accurate identification of pathogens [9,10]. These techniques identify/classify pathogens based on differences in their intrinsic and extrinsic biological properties of bacterial colony. Spectroscopic methods such as Fourier-ransform infrared spectroscopy and Raman [11,12] target chemical constituents, while elastic light-scattering (ELS) sensor [13-15] interrogates morphological characteristics of a bacterial colony. Another sensitive method for pathogen detection is based on mammalian cells-based biosensor that detects live bacteria or active toxins [16,17]. The Bacterial Rapid Detection using Optical scattering Technology (BARDOT) machine is a novel state-of-the-art instrument working on biophysical principles that utilizes the elastic light scatter phenomenon from a bacterial colony to generate signature scatter pattern. BARDOT principle is based on the scanning of bacterial colonies on the surface of agar on the Petri-dishes. It was developed to generate 2-dimensional scatter image of the colonies, then extracting hundreds of linearly independent features to construct an image database. Currently, researchers are building libraries of scatter images of pathogens that could be used by food industries, regulatory agencies and diagnostic or research laboratories as a detection tool to avert outbreaks.
But the major issue with the BARDOT-based sensor is to obtain consistent patterns since many parameters could influence the scatter signatures, like agar concentration, media formulations, and nutrient levels [10,18] . Thus, the goal of this study was to investigate if different preparation methods and storage conditions of agar plate, such as moisture level, air-drying times, storage temperature, and duration of agar plate storage that are generally practiced in a microbiology laboratory, affect the colony scatter signatures. It is known that bacterial growth on solid agar depends on the available nutrients and moisture content [18-22] . Since BARDOT monitors highly sensitive elastic scatter patterns from colonies, slight changes in bacterial growth might affect scatter signatures. Thus, an optimized media preparation and storage conditions must be employed to obtain reproducible data on scatter signatures for various microorganisms. In this study, we examined three different serovars of Shiga-toxin producing E. coli (STEC); E. coli O26, E. coli O111 and E. coli O157 as model bacteria with Sorbitol-MacConkey (SMAC) agar as a model growth media to examine the effect of various environmental/physical parameters on agar media and its resultant effect on scatter patterns of E. colicolonies.
2. Materials and Methods
2.1. Organisms and Growth Conditions
Cultures of E. coli serovar O157:H7 EDL933, O26:H11 HSC27 and O111:H8 HSC27 93-3118 were maintained as frozen stock in 20% sterile glycerol at −80˚C. Cultures were grown in BHI (brain heart infusion) broth for 12 h at 37˚C and 130 rpm in a shaker incubator. Serial decimal dilution of E. coli cultures was made in phosphate buffered saline (PBS, pH 7.4) and spread plated on SMAC agar plates and incubated at 37˚C until the desired colony size (1.1 ± 0.2 mm diameter) was reached, that correspond to fit the 1.0 mm diameter laser beam. Colony diameter was measured using a Leica DMLB microscope (W. Nuhsbaum Inc., USA) with phase contrast setting of 10× magnification. SMAC agar plates with E. coli colonies were scanned by BARDOT and three replicates were averaged for statistical analysis.
2.2. Media
SMAC agar, a recommended medium for the isolation of pathogenic E. coli was prepared as directed by the supplier (Becton Dickinson, USA). Petri dishes with poured SMAC agar (20 ± 2 ml/plate) were allowed to solidify under a laminar-flow bio-safety cabinet (Forma Scientific, USA). Two batches of SMAC agar plates were prepared: Batch-1was used for bacterial growth to be analyzed by BARDOT; and Batch-2 was used for measuring the surface water content using a procedure described in Section 2.5.
2.3. Instrumentation
BARDOT (Advanced Bioimaging Systems, West Lafayette, IN, USA) was used for acquiring the light scattering patterns of bacterial colony (Figure 1). Thisequipment has three principal components: the forward scatterometer, the two-dimensional motorized stage and the colony locator.
It operates with a standardPetri dish with bacterial colony grown on agar media and CCD camera on top that takes images of the petridish to locate individual
 (a)
(a) (b)
(b)
Figure 1. Light scattering sensor based strategy for pathogen detection. (a) BARDOT device; (b) Food extracts or enriched samples are surface plated on agar plate to obtain colonies that are analyzed by BARDOT to obtain unique scatter signatures for specific pathogens.
colonies. The apparatus thus provides a 2-dimensional transmittance map with automatically identified colonycentroid location. Cut-off values for transmission and circularity were chosen to separate single bacterial colonies from doublets (two close colonies). The machine automatically moves the Petri-dish in order to align the colony center with the 635 nm laser. This laser generates the forward scattering pattern as the signature “finger-print” of the colonies [23].
2.4. Image Analysis
The captured laser scatter patterns of bacterial colony were analyzed by the quantitative image processing software. Different features were used for image analysis: the rotation-invariant feature characterized by the Zernike moments of 8; and the Haralick texture feature with a minimum of 1 and maximum of 3 pixels. The performance of the classifier was validated using cross validation [24]. Incase of this experiment, where a single experimental data set is analyzed against the control data set, classification software generates a 2 by 2 confusion matrix (Table 1) which is also called cross-validation (CV) matrix. While CV matrix is more focused on the diagonal terms (true positive and true negative), we define a quantity called Maximum Confusion Factor (MCF) which is the maximum value of either false negative orfalse positive factor (Table 1). A MCF of 10% threshold or higher means that the software makes more than 10% of confusion (or error) between the submitted batches of scatter images and determines that there is more similarity between batches than difference. A MCF lower than 10% means that there are identifiable differences and uniqueness between the submitted batches of images and thus the software classifies the two groups of images as being different with high accuracy. Concisely, higher MCF value represents similar scatter patters and lower MCF value represents distinguishable scatter patterns. More details for the CV matrix are discussed in previous publications [15,23] .
2.5. Surface Water Layer Measurement
Surface water layer (content) on SMAC agar plate was measured by following the published procedure [25,26] where they monitored the influence of agar on the growth of Bacillus cereus. Briefly, an 85 mm diameter filter paper (Whatman, Maidstone, UK) was placed for 1 min on

Table 1. Data interpretation of cross-validation matrixa.
the petri dish after agar solidification. Filter paper weight was recorded before and after absorption of surface water and calculated as μm (absorbed water volume/filter paper area). Surface water layer was measured for SMAC agar plate after subjecting them to short and extreme air-drying conditions under a laminar flow bio-safety cabinet as mentioned in Section 2.6. Subsequently, the BARDOT patterns of three E. coli serovar colonies were acquired after culturing them on air-dried SMAC agar plates.
2.6. Effect of Air-Drying Conditions on Scatter Patterns
To monitor the effect of air-drying conditions on scatter patterns, SMAC agar plates were subjected to short and extreme drying conditions after solidification of the media in a laminar flow bio-safety cabinet. The weight of the Petri dishes was measured before and after pouring the media. For short drying condition, the agar was allowed to air-dry with the plate lid removed, for 0 (control), 10, 20, 30, 40, 50 and 60 min. Similarly, for extreme drying condition, agar plates were allowed to airdry for 0, 3, 5, 7 and 9 h in order to lose 0 (control), 15%, 30%, 45%, and 60% of weight, respectively. A 60% of weight loss in agar plate was previously determined as to be the upper limit for bacterial growth beyond which (>60%), no bacterial growth was observed on the agar (data not shown). During every experiment, before spread plating the bacterial cultures on agar, the average relative humidity (%) and the temperature (˚C) in the room were recorded as 46.8% and 24.8˚C, respectively by using a HMI 31 hygrometer (Vaisala, Finland). Statistical results from scatter images were calculated as described in Section 2.4 after acquiring and analyzing the scatter pattern of E. coli colonies on dried SMAC agar plates.
2.7. Effect of Storage Conditions on Scatter Patterns
To observe the effect of different storage conditions of agar plate on the scatter patterns of E. coli serovars, cultures were spread plated on SMAC agar plates, which were stored for a different period of time: 1, 3, 7, 10, 14, 17, 22, 24 and 30 days. SMAC agar plates were stored in two different conditions in sealed plastic bags: 1) at room temperature (25˚C); 2) at 4˚C in a cold room. SMAC agar plates were weighed at each time point during storage. Weight of empty plate was subtracted to obtain the actual weight of agar. Plastic bags used for storage of SMAC agar plates were sealed with rubber bands. Relative humidity in the laboratory (24.8˚C) and inside the cold room (4˚C) was found to be 46.8% and 72.4%, respectively. The relative humidity and temperature at different conditions of plate storage were constant throughout the study.
3. Results and Discussion
Effects of the preparation and storage conditions of SMAC agar plate on scatter patterns of E. coli serovars O157, O111 and O26 are presented in two groups: 1) effect of air-drying (short and extreme drying durations) agar plates on the scatter patterns; and 2) effect of agar plate storage (at room temperature and 4˚C) on the scatter patterns. The results were tabulated with three distinctive parameters; the condition of drying or storage used in the experiment; the representative BARDOT scatter image obtained under given parameter; and the cross validation matrix obtained with the classification software, where highest off-diagonal values for false positives and false negatives were set as the MCF (Table 1). For all of the data discussing weight loss of agar petri dish, it can be interpreted that weight loss is equivalent to water loss as water evaporates over time. This is the only parameter that can change the weight of the agar in Petri dish over time under different preparation and storage conditions.
3.1. Effect of Agar Drying Conditions on Scatter Patterns
3.1.1. Short Drying Time
As shown in Figure 2, scatter patterns of E. coli O26 and O111 look similar, where as E. coli O157 shows distinct patterns from other two serovars when observed visually. Analysis of the MCF value (Figure 3), for E. coli O26, after 10 min of drying revealed that it is lower than 10%. This means that the software differentiate the patterns of colonies grown on agar that was air-dried for a minimum of 10 min as a unique scatter pattern.
Thus for serovar O26, it can be concluded that, in order to obtain consistent scatter patterns, agar plates should not be air-dried for more than 10 min after solidification. In the case of E. coli O157 serovar, the MCF values were ≥10% until 30 min of drying that means agar plates dried for a maximum of 30 min after solidification will not affect BARDOT scatter patterns of serovar O157. Whereas, scatter patterns of E. coli O111 serovar was least affected from drying agar plates among the three

Figure 2. Diagramshowing the cross-validation (CV) matrix and representative scatter patterns of three serovars of E. coli colonies grown on sorbitol MacConkey (SMAC) agar plates that were subjected to 1 hour of air drying after solidification of agar into Petri dish. In cross-validation matrix the values in diagonal row represent the level of similarity within the submitted batch and anti-diagonal values represent the level of similarity (defined as MCF value in this study) between the two batches of scatter images. A representative scatter image (BARDOT) for each serovar is presented for each time point. Each matrix was computed by comparing the control (T = 0, symbol C) and air-dried plate (T = 10, 30, 50 min, symbols C + 10, C + 30, C + 50). Table 1 provides explanation for each matrix element.
serovars and thus resulting in high MCF values (>10%), even if the agar plates were dried for 60 min (Figure 3). These results suggest that over drying of agar plates for >10 - 20 min on an average after solidification at room temperature under open condition in a bio-safety cabinet may cause variations in the signature scatter pattern of E. coli serovars due to the loss of water content in the agar medium. At this point of study, it cannot be ignored that different serovars of E. coli may have different metabolic activity for the available surface water on agar. Such metabolic differences may have resulted in the differences in the scatter patterns of E. coli O26, O157 and O111 cultured on plates that were air-dried for 10, 30 and 60 min.
From Table 2, it is observed that the surface water layer for the three serovars was relatively constant (28 - 30 µm) for given humidity and temperature within 1 h of short drying time. Calculated surface water layers were around 28 - 30 μm, which was sufficiently higher than the average size of a single E. coli bacterium (0.5 × 2 µm), that should provide enough moisture for the growth of bacteria. The weight loss of agar was recorded over a period of 60 min (Table 2).
As expected, the weight loss of agar in Petri dish increased with increasing drying time. As the drying time increased, MCF generally decreased (<10%) and resulted in differences for scatter patterns of E. coli serovar O26,
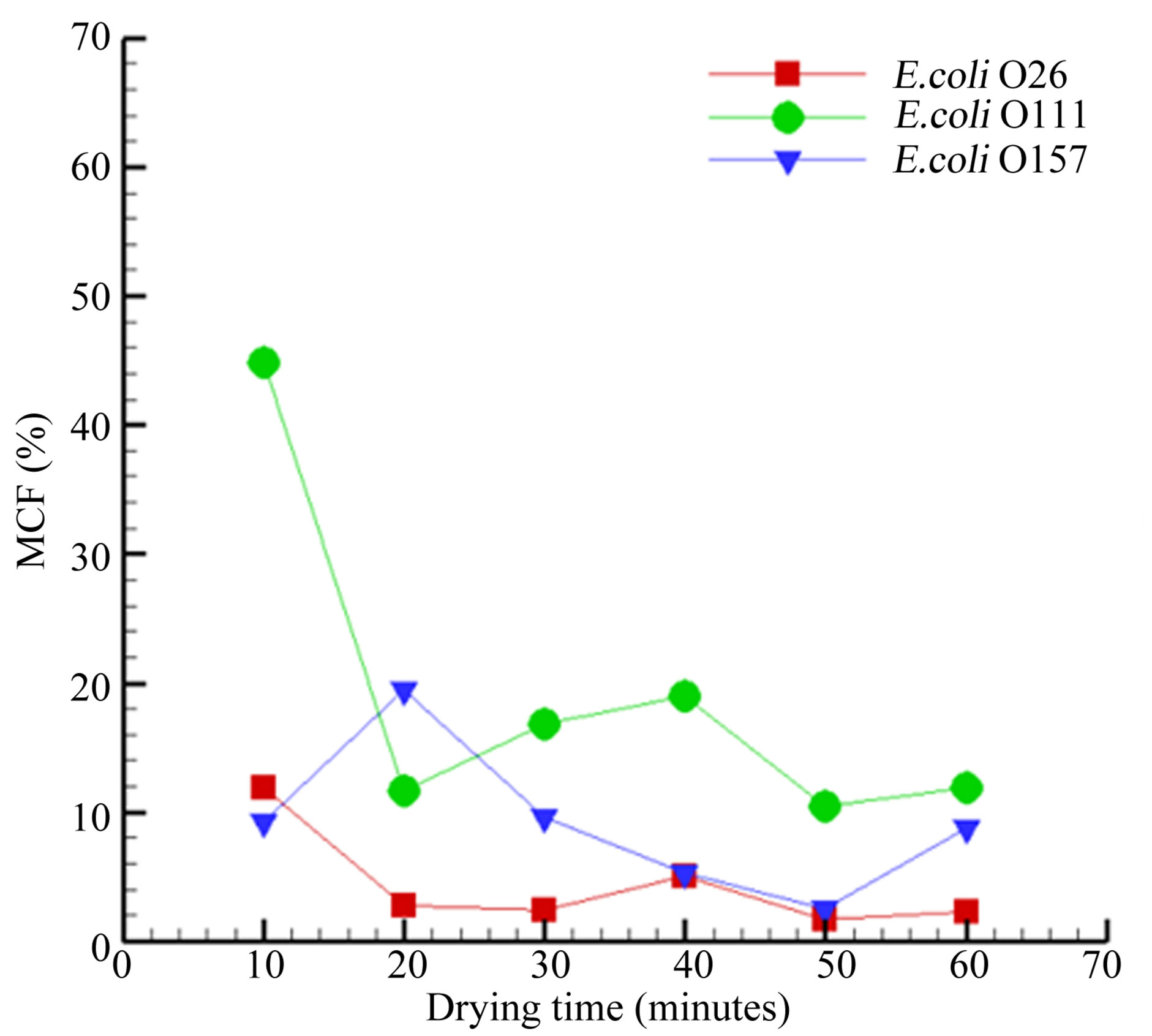
Figure 3. Maximum Confusion Factor (MCF) versus the air-drying time of SMAC agar after solidification followed by growth of three different E. coli serovars. For E. coli O26, between 10 - 20 min of air-drying of the SMAC, the value is lower than 10%. This means that the software can differentiate the patterns of colonies grown on agar that was air-dried for a minimum of 10 min as a unique scatter pattern. Similarly, E. coli O157 patterns were affected when air-drying of plates exceeded 30, whereas E. coli O111 scatter patterns were not affected by air-drying of 60 min.
O157 and O111 at 10, 30 and 60 min of drying time, respectively. This means each serovar was affected differently by short drying time of agar plates (Figure 3). From the results of short drying of SMAC agar plates, it can be concluded that drying agar plates for more than 10 - 20 min resulted in 4% - 8% of weight loss that was sufficient to generate differences in scattering patterns for the same serovar (Table 2, Figures 2 and 3).
3.1.2. Extreme Air-Drying Conditions
We were also intrigued to observe the effect of extreme air-drying conditions (abusive or mal-handling practices) on the scatter pattern. Same analysis was conducted using different air-drying parameters during preparation of the Petri dishes, where the SMAC agar plates were subjected to extreme drying conditions. There was a substantial loss of surface water layer and weight of agar plate during extreme drying condition (Table 3). Results of this study suggest that scatter patterns of serovar O26 were least affected while serovar O157 and O111 were severely affected after growth on plates that were subjected to extreme drying conditions as calculated by MCF values (Figure 4). Data show that for up to 45% of weight loss of agar plate (7 h of drying), BARDOT pattern of E. coli O26 was not affected. However, in the case of E. coli serovar O26, the results of extreme drying (7 h) are contradictory when compared with results of short drying (10 min) and this builds scope for future investigation. In case of E. coli O157 and E. coli O111, MCFs values were lower than 10% during 15% - 45% weight loss of agar plates (Figure 4). However, the high

Table 2. Surface water layer and relative weight loss of the SMAC agar depending on the short drying time (min) after solidification.

Table 3. Surface water layer and relative weight loss of the SMAC agar under extreme drying condition.
MCF value (30% for O111 and 50% for O157) for 60% of agar weight loss is unexpectedly contradictory, and visual inspection confirmed that images are different from the other drying conditions. That shows that human eye and the BARDOT do not analyze the images the same way. The results for 60% of weight loss are not clear and further experiments and analyses are needed. The variations in the findings of short and extreme drying conditions must be related to loss of agar media water content, which was found to be reduced by 2.49% - 7.84% in case of short drying and 15.51% - 61.57% in case of extreme drying conditions. Finding of this study is in perfect analogy with the results of previous investigation [25,26], where 1% and 7% agar was used in media for the growth of Bacillus cereus to mimic short and extreme drying conditions; and observe its effect on available surface water layer. In their study it was observed that Bacillus colony under extreme air-drying condition (7% agar) grew more vertically compared to less drying condition (1% agar), where colony grew more laterally. Such variation in colony profile will impart immense difference in BARDOT scatter patterns based on physical theoretical variations of light forward scattering as mentioned in our previous studies [18,27] . This may be the reason for the irreproducible and incongruent scatter patterns under extreme drying condition, when compared to short drying condition.
3.2. Effect of Storage of Agar Plates on Scatter Patterns
3.2.1. Room Temperature Storage
Storage of SMAC agar plates at the room temperature for5 days revealed low MCF value (<10%) in case of E. coli O26 serovar (Figure 5), which means BARDOT pattern generated on a SMAC agar plate, stored for more than 5 days, started showing differences in scatter pattern for E. coli serovar O26. The high MCF value (>10%) for E. coli O111 and O157 indicates that the storage of agar plates at room temperature for 24 days does not affect the BARDOT pattern (Figure 5). It was also observed that storage of agar plates in sealed plastic bag at room temperature has insignificant effect on the weight loss of agar plates. Here, it can be concluded that the storage of the Petri dishes in sealed plastic bag at room temperature for more than 5 days seems to have an impact on scatter pattern of serovar O26, whereas as it did not affect the pattern of serovar, O157 and O111 on plates when stored for 24 days at room temperature in sealed plastic bag (Figure 5). For up to 24 days of storage of agar plate in sealed plastic bag at room temperature, the surface water layer on the agar was not significantly different from the one of a fresh agar plate prepared at day one (data not presented).

Figure 4. Effect of weight loss of agar on the MCF value for scatter patterns of colonies of E. coli serovars. SMAC agar plates were air-dried for 3, 5, 7 and 9 h after solidification with plate lid open in a laminar flow biosafety cabinet. There was a substantial loss of surface water layer and weight of agar plate during extreme drying condition (see Table 3).

Figure 5. MCF values for scatter patterns of colonies of three E. coli serovars versus the storage time of SMAC agar plates at room temperature for up to 30 days prior to surface plating. For E. coli O26, between 3 - 7 days of storage of SMAC prior to plating resulted in the change in the scattering patterns of colonies, while E. coli O111 scatter patterns remained stable up to 30 days of storage of SMAC plate before plating. Finally, E. coli O157 colony scatter patterns were affected when SMAC plates were used after being stored for 3 - 10 days at room temperature.
3.2.2. Cold Room Storage
By examining data presented in Figures 6 and 7, it can be stated that the MCFs for the three serovars are higher than 10% for most of the time except for E. coli O157 and O26 after 7 or 10 days of storage. Then againafter 14 days of storage, the MCFs are higher than 10%. For the two serovars (O157 and O26), there must be additional factors that may cause changes during this storage period. However, for E. coli O111, the MCFs were all the time higher than 10%, which means that the SMAC agar plates can be used to acquire BARDOT scatter patterns even after one month from the date of preparation, when stored in sealed plastic bag under refrigerated condition.
Figure 7 indicates that certain factor possibly affects the BARDOT scatter pattern on day 7 and day 10 of storage. It does not appear that the surface water layer affected the BARDOT scatter images, because for those days, there was no significant differences on the surface water layer when compared with the others days (data not shown). Results obtained in this study revealed some limitations, which need further investigation: 1) this study was conducted using the three serovars of only one bacterial species grown on single selective medium; 2) weight loss and surface water layer experiments were extremely sensitive and dependent on relative humidity of the laboratory environment, which is influenced by seasonal variations throughout the year; 3) lack of Sensitive instrument and isolated, controlled environment for the experiments. This study has to be expanded to include growth of different bacteria on selective or nonselective media. Application of sensible instrument to measure weight loss, surface water layer and other environmental parameter under isolated system will definitely reduce the level of variations in the experimental setups and measurement. Extreme drying conditions for SMAC agar plate did not result in consistent scatter pattern and it must be emphasized that such extreme conditions of drying is not usual conditions of media preparation in the routine microbiological laboratory setup. Such extreme conditions were inducted in our study to analyze the level of sensitivity and robustness of the BARDOT system.
In future, it would be beneficial to monitor the effect of other experimental variables such as oxygen and nutrient availability on the optical scatter pattern of bacteria, which can also be quantified by determining MCF. Furthermore, the data from this study will help generate a numerical growth model of bacteria under altered environmental conditions. A quantitative MCF value also can be used as one of the parameters to observe the effect of various nutrients, chemicals or antibiotics in media formulation

Figure 6. Diagram representing the cross-validation matrix and representative scatter patterns of the three serovars of E. coli colonies after growth on SMAC plates that were stored in cold room. SMAC plates were subjected to air-drying up to 10 min after solidification and then stored in cold room for up to 30 days. In cross-validation matrix values presented in diagonal rows represent the level of similarity within the submitted batch and the values in anti-diagonal rows represent the level of similarity (defined as MCF value in this study) between the two batches of scatter images. Each matrix were computed by comparing the control (T = 0, symbol C) and stored sample (T = 3, 14, 30 days, symbols C + 3, C + 14, C + 30).
that alter gene expression on optical scatter pattern of foodborne bacterial pathogens.
4. Conclusion
In this study we examined the optimal media (SMAC agar plate) preparation and storage conditions to acquire a reproducible BARDOT scatter patternfor E. coli serovars. Lower MCF values (<10%) for short air-drying period indicate that the agar plates should not be dried after solidification in a laminar flow biosafety cabinet for more than 10 - 20 min. Prolonged air-drying may affect bacterial growth yielding inconsistent scatter patrns. Low MCF values and non-reproducibility of BADORT scatter pattern also suggest that extreme air-drying condition for agar plate should be avoided. This study also underscores the feasibility of preparing and storing agar plates in cold room for 30 days in sealed plastic bag, as it did not affect the colony scatter patterns, whereas storage of agar plates at room temperature does affect the scatter pattern during the course of time for one E. coli serovar. Effect of air-drying and storage conditions are summarized in Figure 8. This finding is extremely important

Figure 7. MCF values for scatter patterns of colonies of E. coli serovars versus the storage time of SMAC agar at cold temperature (4˚C) for up to 30 days prior to plating. For most cases, MCFs are higher than 10% except for E. coli O157 and O26 for the plates that were stored for 7 and 10 days.

Figure 8. Optical scatter patterns of colonies of E. coli grown on solid agar media that were prepared and stored under various conditions.
as it provides guidance for preparation and storage of agar plates in cold room in advance, when BARDOT has to be used in high throughput screening of bacterial pathogens as a contaminant in food samples or for diagnostic purposes on large scale.
5. Acknowledgements
A part of this research was supported through a cooperative agreement with the Agricultural Research Service of the US Department of Agriculture project number 1935- 42000-072-02G and the Center for Food Safety Engineering at Purdue University.
NOTES