Advanced Hierarchical Fuzzy Classification Model Adopting Symbiosis Based DNA-ABC Optimization Algorithm ()
Received 22 January 2016; accepted 20 March 2016; published 23 March 2016

1. Introduction
Good classification allocates given samples to pre-defined classes correctly, and the procedure can be viewed as a type of statistical method. Traditionally, classification is divided into the two stages of training and testing. In the training stage, statistical techniques are employed to obtain classification rules for each feature dimension, while the most common complication in classification is that training samples belonging different categories overlap in some feature dimensions. Pattern recognition is carried out based on the position of each sample spread of each feature dimension in the testing stage. There are suitable statistical methodologies to find the correct class for given samples, such as the Principal Component Analysis, Linear Discrete Analysis methods. Apart from classic statistical methods, expert systems have become the mainstay for classification rate improvement. Additionally, some classification traps and issues need to be addressed to acquire better classification rate.
There are some classification problems, for instance in high-feature space, where some samples overlap in different categories. This leads to mis-categorization and the resulting classification rate may be poor because the training data are misclassified. The problem of partitioning of input features is a big issue while dimensionality is another key obstacle to classification. Therefore, the choice of membership function is considered of paramount importance for fuzzy classification. For these classification problems, it is difficult to use a crisp function to assign samples to the right categories. Generally, some classifiers based on expert systems, i.e. fuzzy or neural networks, are proposed to figure out these issues. For instance, labeled data can be used to establish fuzzy rule-based classification system by GA [1] or SOM tree algorithm [2] . SC-BPN [3] attempts to construct a classifier based on statistical normalization and back propagation. These algorithms are all beneficial based on fuzzy system and neural networks. Briefly, it is not easy to obtain optimal decision performance through traditional fuzzy system or neural networks alone.
In the literature, there are some evolutionary algorithms, such as genetic algorithms (GA) and DNA computing, which improve the classification rate of expert systems based on fuzzy or neural networks. GA includes the operations of reproduction, crossover, and mutation. Data-driven fuzzy classifier (DDFC) [4] utilized co-evolu- tionary methodology is applied to design the fuzzy classifier. The GA-BPN [5] utilized evolutionary algorithm, the FRS-OC-GA [6] based on compact fuzzy rule system with GA, hierarchical fuzzy rule based classification systems with genetic rule selection (HFRBCS) [7] , and Zhou and Khotanzard’s fuzzy-rule-based classifier using genetic algorithms [8] are all proposed to improve the categorization accuracy through evolutionary algorithm.
Similar to genetic algorithms (GA), DNA coding and computing can strengthen the capacity of fuzzy classifier by keeping the elite. Initially, the Hamiltonian path problem was solved using DNA in test tubes in 1995 by Adleman [9] , who was the first person utilizing the DNA for computation. Recently, there is a comparison between the DNA coding and regular coding process by Suang [10] , who applied DNA coding in a GA fuzzy classifier. It was found that DNA coding produces a larger, more diversified population compared to regular coding. Feng [11] enhance the membership function construction by using DNA computing. DNA computing includes the operations of transformation, translocation, enzyme, and virus [12] , which resemble those operations of GA and reinforce it. Therefore, the merit to incorporate the concept of DNA can confer advantages to the next generation.
Learning from the life systems, artificial life systems have been developed to solve the complicated engineering problem in recent decades. Several kinds of population-based algorithms have been utilized for scientific computation. For instance, the fuzzy classification system is improved through ant colony optimization (ACO) based local searcher [13] , feature selection for parameter optimization by using the methodology of fuzzy criterion (FOF) with ACO algorithm [14] , image registration optimization solved by firefly algorithm (FA) [15] , data clustering improved by shuffled frog-leaping algorithm [16] , multi-objective numerical problems solved by the artificial bee colony (ABC) algorithm [17] , and [18] adopting GA and ABC to enhance the fuzzy input feature selection.
Compared to other nature-inspired techniques, ABC has been shown to be very competitive amongst population-based algorithms. The ABC algorithm enhances clustering efficiency and accuracy. Karaboga et al. [19] [20] implement it on three benchmark problems from the UCI machine learning repository and also compared performance versus the PSO algorithm and nine other classification techniques on thirteen databases. In addition to the shortcomings of only using Fuzzy c-means (FCM) are the difficulty of escaping local optima and sensitivity to the initial position chosen, the FCM clustering process combined with the robust ABC searching algorithm may enhance clustering and could be implemented for risk analysis [21] . Apart from the traditional issues of premature convergence and slow convergence traditionally, the global and local searching mechanism of the ABC algorithm enhances the solving of optimization problems. These results seem to satisfy the requirement of clustering. The fuzzy classification flow chart in this study is shown in Figure 1, where symbiosis based DNA- ABC optimization algorithm is utilized to adjust the Hierarchical Fuzzy Classification model.
In the proposed optimization algorithm, as shown in Figure 1, the advantage of the cooperative coevolution mechanism of symbiosis enhances the communication between different populations which results in higher classification rate. In this study, the cooperative coevolution mechanism of symbiosis could benefit artificial bee colonies, and the information exchange between different populations is achieved via hybrid DNA computing, ABC algorithm, and hierarchical based fuzzy classifier. The concept of DNA and the ABC algorithm can be hybridized to realize the cooperative coevolution mechanism of symbiosis in hierarchical fuzzy classification and improve the optimum combination of membership function and the classification rate. The iterative algorithm in ABC added new rules to eliminate unsuitable membership functions, adjust and improve the global optimization capability, and to overcome the “premature” phenomenon.
The proposed symbiosis based modified DNA-ABC optimal algorithm is validated through classifying five benchmark databases: the UCI Pima Indians Diabetes, Glass, Wisconsin Breast Cancer, Wine, and Iris databases. The computer simulation results demonstrate the proposed strategy provides sufficiently high classification rate compared to other models proposed in the literature. The remainder of this paper is structured as follows: Hierarchical fuzzy classification is introduced in Section 2. DNA coding and computing is addressed in Section 3. The idea of artificial bee colony and the details of the symbiosis based algorithm are described in Section 4. The consequence of the experiments and discussions are addressed in Section 5. The conclusion is presented in Section 6.
2. Fuzzy Classification
Over the last few decades, fuzzy systems have been successfully applied to various scientific issues and engineering domains. These fuzzy systems are able to solve difficult problems, exhibit robust behaviors, present linguistic representations of situations, and offer clear advice in regards to classification results. The concept of fuzzy sets was introduced by Zadeh [22] , who proposed using fuzzy logic and fuzzy rules to carry out data processing in the form of linguistic if-then rules. In [8] , Zhou noted for fuzzy system identification and fuzzy parameter optimization, there are three considerations when a fuzzy system is constructed: the selection of feature inputs, feature dimensions partition, and fuzzy rule inferences. Moreover, the fuzzy parameter optimization must consider the associations among the rules and the shape of membership functions.
Fuzzy sets apply fuzzy logic and fuzzy rules to carry out data processing in the form of linguistic if-then rules. The fuzzy classification of this study addresses how to sort several samples into correct categories. It is important to select the correct partition number of membership functions.
Not enough partition number of linguistic values in each attribute causes samples to be classified into the wrong categories. However, increasing the number of linguistic values in each attribute contributes to the calculation time. This increases possibility of failure in databases with much higher dimensional features. The fuzzy
![]()
Figure 1. Symbiosis based DNA-ABC algorithm.
classification in this study can be simply divided into three parts: sample training, fuzzy subsystem generation, and classification unit construction.
2.1. Hierarchical Fuzzy Classification
Triangular membership functions are adopted in this study and an example of fuzzy set with triangular membership functions is shown in Figure 2. Three triangular membership functions are used to uniformly divide each search space into three subsets, where S, M, L partitions denote small, medium, large in Figure 2(a). The SM and ML partitions indicate small medium and medium large in Figure 2(b) and Figure 2(c). Generally speaking, the partition number is equal to the number of triangular membership function.
2.2. Sample Training
It is assumed that there are τ samples for fuzzy classification, and it can be defined as
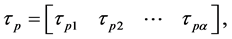 (1)
(1)
where p is the index number of τ samples, and α represents the number of dimensional searching spaces in Equation (1). The linguistic rule of each feature dimension is developed after the membership value 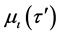 calculated by the following Equation (2).
calculated by the following Equation (2).
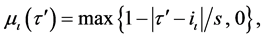 (2)
(2)
where 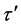 denotes the training sample, s in Equation (2) is the slope of each membership function which equals to 2/(κ − 1), ι is the number of partitions from 1 to κ, and iι is the intercept of each membership function which equals to
denotes the training sample, s in Equation (2) is the slope of each membership function which equals to 2/(κ − 1), ι is the number of partitions from 1 to κ, and iι is the intercept of each membership function which equals to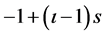 .
.
The linguistic rule (If-Then) enhances the fuzzy classification of numerical input samples in the α-dimen- sional search space of a hierarchical fuzzy sub-system. For ease and convenience, the linguistic rule with three inputs adopted in this study to solve the classification problem is Equation (3),
![]() (a)
(a)![]()
![]() (b) (c)
(b) (c)
Figure 2. Membership functions.
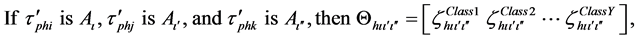 (3)
(3)
where h is the number of hierarchical fuzzy subsystems equal to α/2. The subscripts hi, hj and hk are the searching spaces of h in α. The subscripts ι, ι', and ι" are the number of fuzzy subsets in the h of α. Aι, 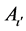 , and
, and 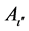 are the antecedent fuzzy subsets. ζ represents the corresponding rule confidence grade, which means the higher value of ζ the higher probability that this rule will assign the training sample to the category Y, and Θ is consequents for each fuzzy rule.
are the antecedent fuzzy subsets. ζ represents the corresponding rule confidence grade, which means the higher value of ζ the higher probability that this rule will assign the training sample to the category Y, and Θ is consequents for each fuzzy rule.
2.3. Generation of the Fuzzy Subsystem
The entire training sample membership function sum for each fuzzy subset Aι, 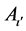 , and
, and 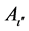 are calculated with equations (4) and (5) at the beginning of the process of fuzzy subsystem generation. The confidence grade of each fuzzy rule for each class are assigned,
are calculated with equations (4) and (5) at the beginning of the process of fuzzy subsystem generation. The confidence grade of each fuzzy rule for each class are assigned,
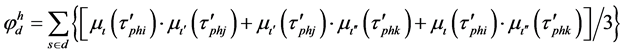 (4)
(4)
![]() (5)
(5)
where ![]() is the membership function of the antecedent fuzzy subset A, and class order
is the membership function of the antecedent fuzzy subset A, and class order![]() . After training, applying test datum into the established if-then rule allows the inferred strength of three input samples to be calculated as Equation (6),
. After training, applying test datum into the established if-then rule allows the inferred strength of three input samples to be calculated as Equation (6),
![]() (6)
(6)
where the inferred strength ![]() is the probability of classifying each sample to each category, and “
is the probability of classifying each sample to each category, and “![]() . L represents the number of rules inferred from the input of the subsystem, which is not fixed and varies according
. L represents the number of rules inferred from the input of the subsystem, which is not fixed and varies according
to the inference result of each subsystem. According to Equation (6), the inferred strength ![]() is the product of the testing sample’s membership function and the resulting
is the product of the testing sample’s membership function and the resulting ![]() from the training samples of each rule.
from the training samples of each rule.
2.4. Classification Unit Construction
After getting the inferred strength ![]() of each subsystem, the following equation calculates all subsystem to classify the testing datum:
of each subsystem, the following equation calculates all subsystem to classify the testing datum:
![]() (7)
(7)
![]() (8)
(8)
![]() (9)
(9)
where ΨH, confidence level, is the sum of each subsystem for every class, as calculated by Equation (7). The classification result Gr is determined by using Equation (8) and Equation (9) obtains the maximum G to determine which class the testing sample should belong to.
3. DNA Computing
The chromosome population and generation are defined initially, and the objective function should be defined before DNA computing, which can be expressed as
![]() (10)
(10)
where the true positive pattern mean the event that the test sample makes a positive prediction and the subject has a positive result after classification, and the number of all patterns equals to the number of test samples.
To construct several chromosomes representing the number of membership functions and the related shift in nucleotides, the objective function is the classification rate equaling to true positive patterns divided by all patterns, which is expressed as Equation (10), and the number of population is related to the input feature dimensions, i.e. the higher the number of input feature dimensions, the larger the population. All the DNA chromosomes are adjusted through the objective function and DNA computing, e.g. duplication, mutation, etc. The pseudo-code of canonical DNA computing algorithm used in this study is given as Pseudocode 1 .
In general, constructing the chromosome by DNA coding simplifies the coding process. Adenine (A), Guanine (G), Cytosine (C), and Thymine (T) are the four basic elements of biological DNA, while codons is a combination with a triplet code of nucleotide bases, such as ATG or CAA. The DNA index of codons is listed in Table 1.
![]()
Pseudocode 1. Pseudocode of canonical DNA computing.
a. Codons―Corresponding number.
In this study, there are sixty-four three-letter combinations. The corresponding number of each codons is used in DNA coding, and it is defined as number 0 to number 63, which is from the top left block to the bottom right block. Inspired by Jan [11] , who utilized DNA coding to construct structures of P, PD, PI and PID in a PID controller, the crossover, mutation, enzyme, and virus operations is applied to obtain the fittest structure. The chromosome structure used in this study is changed dynamically by defining each chromosome as two partitions, one is the membership function and the other is related to the shift nucleotide as shown in Figure 3.
In Figure 3(a), the first two three-letter nucleotides in front of each DNA chromosome are used to describe the membership function of the fuzzy subsystems, which can then be transferred to a twelve-digit binary code to encode the chromosome. The residual part of the DNA chromosome is the shift nucleotide, which contains the corresponding number of fuzzy subsystem shifts. Therefore, the initial first two three-letter nucleotides in front of each DNA chromosome are both GGG, and the numbers of the membership function of each fuzzy subsystem is twelve as shown in Figure 3(b). Furthermore, an example of the first two three-letter nucleotides is shown in Figure 4.
In this example, the two DNA nucleotides at the front of the chromosome are ACU and CUC. The ACU and CUC are strings for the decimal numbers 36 and 17, respectively, and when transformed to binary code they become 100100 and 010001, respectively. Through DNA coding, there are four membership partitions, M1, M4,
M8, and M12, and the other membership partitions are eliminated. The four partitions are then changed to![]() ,
, ![]() ,
, ![]() , and
, and ![]() and the available shift distance Δ between each partition is shown as Figure 5.
and the available shift distance Δ between each partition is shown as Figure 5.
![]()
![]() (a) (b)
(a) (b)
Figure 4. An example of the first two three-letter nucleotides of DNA chromosome.
![]()
Figure 5. Shift distance between each triangular peak calculation.
The distances between each triangular peak are calculated after each membership partition has already been established, as shown in Figure 5, and the membership function can only shift half of the differential distance in
the left or right directions. For example, ![]() is the differential distance between the two peaks of the
is the differential distance between the two peaks of the ![]() and
and ![]() membership functions. Equation (10) reveals the rule of shift direction and distance.
membership functions. Equation (10) reveals the rule of shift direction and distance.
![]() (11)
(11)
Therefore, if the number transferred from the DNA nucleotide according to Table 1 is less than or equal to 32, then the peak of the membership function shifts proportionally to the left as in Equation (11). In the same way, the peak of the membership function shifts to the right if the number is greater than 32. The example is given as Figure 6.
In Figure 6(a), there are four additional nucleotides at the residual part of the chromosome. The two red codons define the ![]() and
and ![]() membership function peak shift distance. The two orange codons define the
membership function peak shift distance. The two orange codons define the ![]() and
and ![]() membership function peak shift distance. These numbers are 32, 53, 21, and 33, as seen in Table 1, and the DNA coding combinations used in the chromosome are AUU, GCC, CCC, and AUC, respectively. The peaks of M2" partition and M3" partition need to shift right after adopting Equation (11), which is shown as Figure 6(b). Additionally, DNA computing includes crossover and mutation process are shown in Figure 7.
membership function peak shift distance. These numbers are 32, 53, 21, and 33, as seen in Table 1, and the DNA coding combinations used in the chromosome are AUU, GCC, CCC, and AUC, respectively. The peaks of M2" partition and M3" partition need to shift right after adopting Equation (11), which is shown as Figure 6(b). Additionally, DNA computing includes crossover and mutation process are shown in Figure 7.
In Figure 7(a) and Figure 7(b), two nucleotides from the chromosome are randomly chosen and then exchanged in the crossover process, and nucleotides are randomly chosen and changed when a chromosome is selected for mutation. These two kinds of DNA computing are similar, which only replace the DNA nucleotide by another one, but not change the length of DNA nucleotide. In this perspective, the virus and enzyme computing is shown in Figure 8.
The virus computing shown in Figure 8(a) is to eliminate the shift nucleotide and decrease the number of membership function peak after transforming from DNA codon to binary code, while the shift nucleotide is added and the number of membership function peaks increase after transforming from DNA codon to binary code in the enzyme computing as shown in Figure 8(b). As a matter of fact, the mutation rate is set at
0.1 in
this
![]()
![]() (a) (b)
(a) (b)
Figure 6. An example of the shift nucleotides of DNA chromosome.
![]()
![]() (a) (b)
(a) (b)
Figure 7. An example of DNA chromosome computing.
![]()
![]() (a) (b)
(a) (b)
Figure 8. An example of DNA chromosome mutation computing.
study to avoid the distractions of result. After DNA computing, to maintain the superiority of the population, the fitness value is calculated based on the objective function and wheel duplication is adopted for the next population.
4. The Symbiotic Based Artificial Bee Colony
The superiority of the artificial bee colony (ABC) algorithm in both unconstrained optimization problems and constrained optimization problems has been proven [23] [24] . The behaviour of a foraging artificial bee colony inspired the meta-heuristic algorithm of ABC. The original idea was proposed by Karaboga to optimize real parameter optimization in 2005 [25] . Karaboga also adopted Deb’s heuristic selection rules [24] to substitute for the greedy selection of the ABC algorithm and applied it to thirteen well-known constrained optimization problems. The comparative results show the ABC algorithm is superior than the other two population based algorithms (differential evolution (DE) and particle swarm optimization (PSO)) for constrained optimization problems. Consequently, using hybrid artificial bee colony and genetic algorithm (GA) on the constrained function is more efficient than using GA only [26] . The real-parameter optimization through the modified ABC algorithm also acquires suitable results on hybrid functions compared to other state-of-the-art algorithms [27] . Therefore, the ABC algorithm is adopted in this study.
The word “symbiosis” was defined by Professor Anton de Bary in 1879 as “the living together of unlike organisms” [28] . This concept is implemented and expanded in several algorithms. As described in [29] , they enhanced a particle swarm to solve six non-linear function minimization problems based on the symbiosis concept. The competitive version of multi-swarm cooperative particle swarm optimizer (COM-MCPSO) and the collaborative version of MCPSO (COL-MCPSO) are proposed according to the co-evolutionary relationships between master swarm and slave swarms. Inspired by [30] , a constrained multi-objective optimization problem was solved using the concept of symbiosis with ABC. The authors also discussed the characteristics of symbiosis between dissimilar populations and the characteristics of cooperation. The symbiotic structure is shown as Figure 9, where each row represents one artificial bee colony and each column represents different feature dimensions. Information between different artificial bee colonies is translated through this symbiotic mechanism. The symbiotic structure considers both social evolution and cooperative coevolution through individual exchange and information sharing.
Three artificial bee groups are simulated in an ABC algorithm, the employed bees, onlooker bees and scout bees. The ABC algorithm attempts to balance exploration and exploitation. The search processes of the employed bees, the onlooker bees and the scout bees are subjected to repeat the foraging cycle until a predetermined number of cycles is reached. In particular, the boundary value of forage position and the maximum number of searching cycle affects the optimization process significantly. A larger number will slow down the performance, and too small a value cannot meet the requirements which are addressed in [31] . The employed bees and the onlooker bees implement the local search while the onlooker bees and the scout bees carry out global search methods.
Typically, the number of the employed bees and the onlooker bees are assumed to be equal to the number of forage but half the number of the colony. The number of scout bees is equal to the number of employed bees who abandoned the food source. The chance value for an onlooker bee selects nourishment source calculated as Equation (12) described in [25] .
![]() (12)
(12)
where j in Equation (12) indicates the nourishment source, where the associated Cj is the chance value of nourishment source selected by an onlooker bee, sutj is the nectar amount measured by the employed bees in the j-th nourishment source, and FN is the nourishment source number. The local search strategy for finding the new possible solution of the original ABC optimization algorithm could be seen as Equation (13), which is also described in [25] .
![]() (13)
(13)
where υef in Equation (13) is the new candidate nourishment source, while χef and χef represent various old nourishment source positions where e and f are integers between [1, FN] and [1, α], respectively. α is the dimension number needed to be optimized. The difference between χef and χgf represents the distance from one nourishment source to another. γef influences the distance of a neighbour nourishment source position around χef, and is a random number between [−1, 1]. The constraint of Equation (13) is g must be different from e.
4.1. Canonical Artificial Bee Colony
The initial positions of nourishment sources are generated randomly. Each employed bee learns the nectar amount of these local nourishment sources, compares the nectar information of each previous and new nourishment source. It memorizes the new position and forgets the old one if its nectar is higher than that of the previous one, and then it updates and shares the latest quality information through waggle dances to the onlooker bees waiting in the nest. An onlooker bee evaluates the nectar amount information taken from all employed bees and chooses a nourishment source with a probability related to its nectar amount. Nourishment sources will be abandoned and replaced with a new nourishment sources by the scout bees. In other words, if the nourishment source can not be improved further to obtain the adequate quality of nectar and then it will be abandoned. The expression of the canonical ABC algorithm is given as Pseudocode 2 .
However, the insufficiency of accelerating convergence speed and escaping local optima are still issues for search equations is addressed in [32] . The idea to integrate population-based algorithms with evolution-based algorithms is inspired by the characteristic of cooperative coevolution from symbiosis.
4.2. Symbiosis Based Modified DNA-Artificial Bee Colony
The initial chromosomes with different feature dimensions are generated randomly and coded by the codons table. The following is the repeat cycle of employed bee: it acquires the fitness, which is evaluated by fuzzy classification, of these chromosomes, and compares the fitness information of each previous and new chromosomes, it memorizes the new and forgets the old if its fitness is higher than that of previous one, it updates the latest quality information and shares it to the onlooker bees. An onlooker bee evaluates the fitness information taken from all employed bees and chooses a chromosome with a probability related to its fitness. The chromosome will be abandoned and replaced with a new chromosome during the scout bees repeat cycle. After that, chromosomes with different dimensions translate their information by DNA computation after each cycle to generate better chromosomes.
The Equation (14) and Equation (15) shown as below are the objective functions used for classification.
![]() (14)
(14)
![]()
Pseudocode 2. Pseudocode of canonical ABC algorithm.
![]() is the value obtained through canonical ABC algorithm for each bee colony. The classification rate,
is the value obtained through canonical ABC algorithm for each bee colony. The classification rate,
which is the higher the better, plays an important role in classification. Additionally, number of rule is another key factor needed to be concerned in classification.
![]() (15)
(15)
where nr in Equation (14) means the number of rule. The number of rule has been defined when the DNA coding is applied in each dimension, and the consequent will be understand more easily when the number of rule is small.
Therefore, the value of ![]() should be directly proportional to classification rate, and inversely proportional
should be directly proportional to classification rate, and inversely proportional
to the rule number. Where cr(%) in Equation (14) and (15) is the classification rate, which is equal to true positive patterns divided by all patterns and can be expressed as Equation (16):
![]() . (16)
. (16)
The number of rule and the classification rate will decide the algorithm success or not, which is the main reason should be concerned meanwhile. Therefore, the Equation (17), which is combined the characters of Equation (14) and Equation (15), is the objective function used in this study.
![]() . (17)
. (17)
The expression of the symbiosis based modified DNA-ABC optimization algorithm is given as Pseudocode 3 . Take database which is containing 4 dimensions and 3 membership function partition in each dimension for instance. There are 8 candidate DNA chromosomes in each dimension and 32 candidate DNA coding chromosome totally in the beginning, and 144 rules will be trained in symbiosis based DNA-ABC optimization algorithm through hierarchical fuzzy classification model. Each bee on behalf of one rule in the proposed algorithm, and then setting up the parameter of forage, limit, and the maximum repeat cycle 100, 100, and 100, to get the local optimum candidate DNA chromosome, apply objective function with DNA computing 10 times to get the global optimum of candidate DNA chromosome finally.
![]()
Pseudocode 3. Pseudocode of symbiosis based modified DNA-ABC algorithm.
The cooperative coevolution mechanism of symbiosis benefits the different swarms, and the information exchange between different populations is achieved by hybrid DNA computation and hierarchical based fuzzy classifier.
There is more than one feature dimension in one data set, and the different feature dimensions in each data set could be viewed as different artificial bee colonies. With the convenience of the chromosome structure for each artificial bee colony, an adequate solution in each feature dimension could be obtained by the canonical artificial bee colony algorithm, while a suitable combination solution of different feature dimensions is acquired through the symbiosis based artificial bee colonies algorithm. The optimum classification rate is achieved through the cooperative coevolution mechanism of symbiosis.
5. Experimental Results and Discussion
It is essential to test classifier performance by implementing and testing on adequate databases. For instance, the ORL and Yale face databases could be the most popular to test a human face image based classifier, etc. In this study, number based databases are selected for classification. There are several famous databases collected by the University of California at Irvine (UCI). Generally, the five most commonly used databases in the literature for classification are the Pima Indians Diabetes (PID), Glass, Wisconsin Breast Cancer (WBC), Wine, and Iris databases. Details of these databases are shown in Table 2, where the sample numbers are ordered from large to small.
It is difficult to evaluate the classification rate of data set only through the number of samples or classes. For instance, the Iris data set contains 3 classes, with 50 samples for each class, while Wine data set also contains 3 classes, but 59, 71, 48 samples for each respective class. The number of samples and classes are similar; however, the features are different. For example, the Iris data set consists of four features: sepal length in cm, sepal width in cm, petal length in cm, and petal width in cm, while the Wine data set consists of thirteen features: Alcohol, Malic acid, Ash, Alcalinity of ash, Magnesium, Total phenols, Flavanoids, Nonflavanoid phenols, Proanthocyanins, Color intensity, Hue, OD280/OD315 of diluted wines, and Proline.
From previous experience in [33] - [35] , the classification rate of the Iris dataset is lower than the Wine data set. The reason might be that the optimum classification rate occurred only when the training samples contain
sufficient information. Therefore, five different databases are used to demonstrate the reliability of the proposed hierarchical fuzzy classifier.
The two main approaches utilizing to evaluate the performance of the symbiosis based modified DNA-ABC algorithm are using all data in each database for training approach and the leave-one-out approach where one sample is removed from the dataset as the testing sample, and repeat until all of the samples have been used for testing. The simulation results obtained using hierarchical fuzzy classifier only is shown in Table 3 and Table 4. The approach using all data to train and test in the original model is shown in Table 3. The approach using leave-one-out is shown in Table 4.
Comparing Table 3 and Table 4, the classification rate and rule numbers are the same when classifying the Iris database. However, the simulation result of the other four databases in Table 3 is better than Table 4. The reason is that some data may not supply suitable information for training which lowers the classification rate. The symbiosis based modified DNA-ABC algorithm setting is implemented as follows: The parameter of forage, limit, and the maximum repeat cycle is setting up 100, 100, and 100, which are the same as [36] , and apply objective function with DNA computing 10 times. The membership function adjusted by symbiosis based DNA- ABC algorithm, and the consequence is shown in Table 5 and Table 6.
In Table 5 and Table 6, the number of partition in each feature space is setting to twelve of all databases at the beginning of the process. The status “Before DNA-ABC tuning” means the membership function not be tuned yet, and “After DNA-ABC tuning” means that the number of rules is reduced by using symbiosis based DNA-ABC algorithm to adjust the membership functions to minimize the number of if-then rules in the whole rule base. Obviously, the classification rates and rule numbers are all improved after turning. The classification rate is compared with other expert systems in the literature using fuzzy and neural networks in Table 7.
In Table 7, Roubos’ algorithm [1] , TSFR [2] , SC-BPN [3] , DDFC [4] , GA-BPN [5] , FRS-OC-GA [6] , HFRBCS [7] , Zhou [8] ’s algorithm, and FOF [14] , populate the following columns. Compared with other classification experiments, our method gives the best classification rate for three of the five databases. The symbiosis based modified DNA-ABC algorithm efficiently enhances the hierarchical fuzzy classification system performance through the cooperative coevolution mechanism. Note a higher partition number in the membership function acquired from this algorithm will slow down the speed of classification.
6. Conclusion
A symbiosis based modified DNA-ABC optimization algorithm for hierarchical fuzzy classification is proposed in this study. It combines the advantage of the DNA coding chromosome structure and DNA computing to enhance communication between different swarms. It overcomes dimensional and overlapping issues and also obtains the optimum classification rate through suitable membership function extraction. The procedure is divided into four main parts. First, membership functions (chromosome) are extracted by DNA coding and evaluated through fuzzy classification. Second, new chromosome (food source) generation and the fittest chromosome are retained in each feature dimension during the employed bee repeat cycle by evaluating the fitness function of each chromosome. Third, the probability value of each chromosome is calculated for each onlooker bee in each dimension during the onlooker bee repeat cycle. Last, the abandon chromosome is replaced by scout bees in different feature dimensions, memorize the optimum chromosome, and implement DNA computing during the scout bee repeat cycle. Traditionally, the ABC algorithm is mainly applied towards constrained and unconstrained problems.
![]()
Table 3. Classification rate obtained by only using hierarchical fuzzy classifier with all samples for training and testing.
b. Classification Rate(%)/Rule Number.
![]()
Table 4. Classification rate obtained by only using hierarchical fuzzy classifier with leave-one-out approach.
![]()
Table 5. Classification rate simulation results with modified DNA-ABC operator with all samples for training and testing.
c. Classification Rate (%)/Rule Number.
![]()
Table 6. Classification rate simulation results with modified DNA-ABC operator with leave-one-out approach.
![]()
Table 7. Hierarchical fuzzy rule based evolutionary programming comparison.
d. Not presented.
In this research, it is integrated with DNA concepts and applied to fuzzy classification. From the best of our knowledge, this approach was not used previously. Simulation results illustrate that the proposed symbiosis based modified DNA-ABC optimization algorithm for hierarchical fuzzy classification can offer the best classification rate in three of five databases. However, calculation time is required to achieve higher system performance, which is a known disadvantage of the canonical ABC algorithm.
NOTES
![]()
*Corresponding author.