Ubiquitin C gene: Structure, function, and transcriptional regulation ()
1. INTRODUCTION
Ubiquitin (Ub) is a highly conserved 76 amino acid protein which can be conjugated to other proteins via an isopeptide linkage between the carboxy-terminal glycine residue of ubiquitin and the ε-amino group of a lysine within the target protein or it can make an isopeptide bond with a lysine in another moiety of ubiquitin to form Ub chains. This post-translational modification is known to play a central role in the regulation of various cellular processes: the number and placement of Ub molecules added to a protein help to determine its fate. The effects of ubiquitination of a target protein range from the well-known protein degradation through the 26S proteasome, to protein trafficking, cell-cycle regulation, DNA repair, apoptosis and signal transduction [1-6]. Ubiquitin is one of the most abundant protein in eukaryotic cells, constituting ~0.1% - 5% of total proteins but, despite its pervasive use and large number of substrates which are ubiquitinated in a cell, Ub does not seem to be produced in excess, rather the free pool of Ub is maintained at an adequate level depending on cell conditions [7-9]. Ubiquitin homeostasis is maintained through different mechanisms: recycling of ubiquitin chains by deubiquitinating enzymes (DUBs) and de novo synthesis (“Figure 1”) [10].
2. UBIQUITIN GENES
Ubiquitin is encoded in the genome by a family of loci. Two of these genes, UBA52 and RPS27a code for a single copy of ubiquitin fused to the ribosomal proteins L40 and S27a whereas UbB and UbC are polyubiquitin genes and encode 3 and 9 head-to-tail repeats of ubiquitin, respectively These genes were first characterized by Wiborg et al. in 1985 [11] using Northern blotting analysis of poly(A)-containing RNA from different tissues, but only few years later they were localized in the genome and fully characterized [12-15]. Peculiarity of Ub genes is the process they undergo to obtain free ubiquitin monomers. The first hypothesis was mRNA processing, but it was ruled out since polyubiquitin and ribosomalfusion transcripts displayed molecular weights on northern blot corresponding to the molecular weights predicted from their genes [11]. In 1989, Monia et al. expressed UBA52 and RPS27a in Saccharomyces cerevisiae and demonstrated that the mechanism responsible for generating the free ubiquitin monomer in eukaryotic cells was a proteolytic process involving specific enzymes [16]. In the same year, the first DUBs were identified, cloned and purified based on the ability to hydrolyze

Figure 1. Main processes contributing to free ubiquitin pool. (1) Fusion products encoded by ubiquitin genes are cotranslationally processed by DUBs, giving free ubiquitin monomers; (2) Editing of mono-, multimonoand poly-ubiquitin tags by DUBs, to modulate the signal outcome; (3) Recycling of polyubiquitin chains after target protein degradation through the proteasome, by DUBs; (4) Protein rescue from 26S proteasome degradation through the removal of polyubiquitin tag, by DUBs. In (2) and (4), the reverse process follows the 3-step enzymatic cascade (activation, conjugation, ligase) for ubiquitin binding to target substrate.
esters or fusion proteins containing ubiquitin [17-19].
3. UBC
Ubiquitin C is a polyubiquitin gene, located on chromosome band 12q24.3 [15]. Its DNA sequence is 2596 bp long and it contains a first untranslated exon (63 bp), an intron (812 bp) and 9 tandemly repeated ubiquitin moieties. UbC gene, together with UbB, was first described as a stress inducible gene, upregulated upon different cell treatments as well as in some pathological conditions. Polyubiquitin gene response to heat shock was first described in chicken embryo fibroblasts by Bond et al. in 1985 [20]. Different cell lines and microorganisms were, from then on, tested for ubiquitin upregulation after heat shock and a great bulk of literature is now available [21-23]. UbC gene was also found to be the most responsive gene to UV irradiation [24,25], oxidative stress [26], translational impairment through cycloheximide and the amino acid analogue canavanine [27,28]. Taken together, these evidences categorize UbC as a stressresponsive gene, at the front line against external treats. Even do UbC together with UbB is thought to only supplement the constitutive monoubiquitin genes (UBA52 and RPS27a) in maintaining cellular ubiquitin homeostasis, there is evidence of the basal importance of UbC in dealing with this role: mice harbouring a targeted disruption of the UbC gene (UbC−/−) die between embryonic days 12.5 and 14.5. Moreover, mice embryonic fibroblasts (MEFs) from UbC−/− embryos exhibit reduced growth rates, premature senescence, increased apoptosis and delayed cell-cycle progression and drastically reduced size of liver mass [29]. The midgestation embryonic lethality, the liver development phenotypes and the delayed cell-cycle progression were partially rescued by providing extra genomic copies of Ub. This suggests that the whole phenotype is likely the consequence of Ub deficiency. Steady-state Ub levels were quantified and found to be the 40% in UbC−/− MEFs, compared to the wild type. UbB transcript was significantly increased in UbC−/− mice, but this increase was not enough to compensate for the loss of UbC. Thus, it is possible to state that UbC contributes significantly to the maintenance of steady-state total Ub reservoirs in the cells.
4. UBC TRANSCRIPTIONAL REGULATION
Although the functional importance of UbC has been widely described, the mechanisms responsible for its regulation have not been fully elucidated. The first group trying to reveal the mechanisms involved in UbC transcriptional regulation was led by Nenoi [30]. In 1996 Nenoi et al. putatively identified an intron sequence and a transcription start site, later confirmed by RT-PCR and primer extension experiments, willing to characterize an upstream control region. They also find putative transcription factor binding sites such as: HSF, NF-kB, AP-1 (c-jun), NF-IL6, Sp1 as well as a TATA box sequence. Few years later, in 2002, the first, real molecular study on UbC promoter was carried out by Marinovic et al. [31]. Starting from the evidence that in catabolic conditions, glucocorticoids raise polyubiquitin mRNAs as well as some proteasome subunits [32,33], they explored the molecular mechanisms that enhance transcription of UbC gene. Marinovic et al. provide in vitro and in vivo evidences that UbC induction in rats with catabolic pathologies, is due to an activation of the MAPK signalling pathway which at the end leads to a direct Sp1 binding to promoter target sequences. They also present evidences of Sp1 importance in basal UbC transcription [31]. This mechanism was then found to be cell-specific, influencing UbC expression only in muscle or myeloma cells [34,35]. Hepatocytes, cardiomyocytes, kidney and colon cells didn’t respond to dexamethasone with an increase in UbC expression. UbC promoter was also studied in a model of skin carcinogenesis in mice. Kim et al. found a DR-1 element (PPARβ responsive element) in the promoter sequence of UbC gene which, after hyperplastic treatment with TPA, binds in vivo PPARβ. This leads to an attenuation in skin carcinogenesis [36]. These research papers shed some light on the mechanism(s) involved in UbC upregulation following different treatment or cellular conditions, but what happens in basal physiological conditions remains unknown. The first group addressing this problem, trying to identify the cis-trans regulatory elements that orchestrate UbC basal transcription was led by Bianchi [37]. Starting from the evidences that in eukaryotic expression vectors, introns are inserted in order to enhance transgene expression and that a process called intron-mediated enhancement (IME) was found to regulate plant polyubiquitin genes [38], they wondered if a similar mechanism could be found also in human UbC gene. They cloned three serial 5’-deletion fragments of the upstream region of UbC gene in a promoter-less luciferase expression vector. The longest region cloned spanned from −916 to +878 (with respect to the TSS) and contained a portion of the 5′ flanking promoter sequence, the 63-bp exon 1 and the 812-bp intron of the 5′-UTR of the UbC gene. To investigate the importance of the unique intron, they also removed it from the reporter vectors and transfected them, together with the intron-containing constructs, into three different cell lines (HeLa, SiHa and U2OS). The results, summarized in “Figure 2”, show that the minimal promoter region, required for maximal activity is located within the −371/+878 nucleotide sequence, suggesting that the −916/−371 portion doesn’t contain crucial cis-elements for promoter activity. The removal of the intron showed instead a drastic drop of luciferase expression, meaning that the unique intron is needed for maximal promoter activity. Bioinformatic analysis, supported by EMSA and supershift assay, revealed multiple intronic
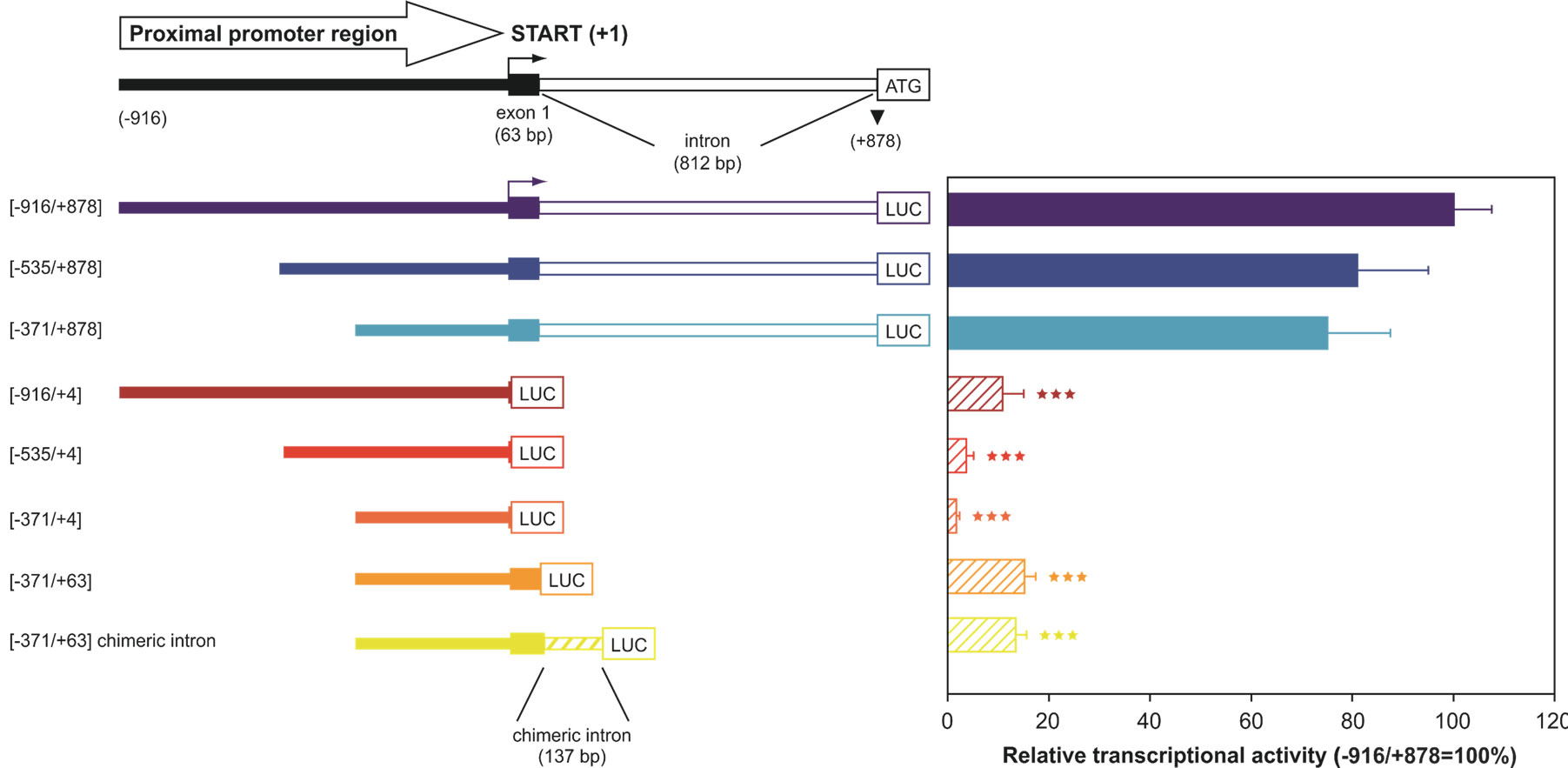
Figure 2. 5’- and 3’-deletion fragments of the UbC promoter, driving luciferase expression. The upper scheme represents the proximal promoter region cloned (nt −916/+878), with indication of the transcription start site (+1, marked by an arrow), exon and intron length. The boundaries of the UbC promoter regions (intron-bearing and intron-less) cloned into the pGL3-basic vector are indicated. The chimeric intron bearing construct is also depicted. Luciferase transcriptional activity measured in HeLa cells transiently transfected with the showed reporter constructs is reported in the histogram. All values were referred to that of the larger construct (−916/+878), which was set arbitrarily as 100 (***, p < 0.001 versus −916/+878 construct).
binding sites, for three ubiquitously expressed transcription factors: Sp1, Sp3 and YY1. Cotransfection experiments, using the construct −371/+878, harboring the intron, and the respective intron-less construct with an Sp1 or Sp3 expression vector showed, for both the reporter constructs, a statistically significant raise in luciferase expression, meaning that these two factors may be involved, in vivo, in UbC expression. Unexpectedly, mutagenesis of Sp1/Sp3 intronic binding sites didn’t result in a drop in luciferase transcripts, indicating that, despite the in vitro evidences (EMSA and supershift), they don’t have an in vivo importance in promoting UbC expression [39]. This is not in contrast with the previous evidence that transfection of Sp1/Sp3 raises the expression of the intron-retaining and intron-less constructs because the enhancement observed could be due to Sp1/Sp3 interaction with the upstream binding sites (contained in both constructs), previously identified by Marinovic et al. [31]. On the other hand, when the two YY1 intronic binding sites were mutated, a statistically significant drop in luciferase activity was observed [39]. A wide number of in vitro (EMSA, supershift) and in vivo (ChIP, mutagenesis, transfection and cotransfection with siRNA) experiments were provided by this group in support to YY1 role in UbC expression. This paper also gives a new insight in UbC regulation, thanks to the study on splicing. A new emerging role for YY1 was indeed found: mutations of its binding sites or a specific siRNA anti-YY1 impaired UbC intron splicing and RNA immunoprecipitation experiments show this notably known DNA binding factor, also bound to UbC mRNA.
5. CONCLUSIONS
Ubiquitin role within the cell is very complex and unique. Its functions extend from the well-known proteasomedependent proteolysis to cell signalling, transcription, apoptosis and so on. This feature is fully reflected in its peculiar gene structure and translational mechanisms. The molecular pathway that goes from mRNA to free ubiquitin monomers is indeed very complex and, based on the above statements, it is not surprising that every step is fine-tuned in order to maintain an adequate ubiquitin pool to face different cell challenges. This review is aimed at giving an overview of what is known and what is missing in the mechanisms regulating ubiquitin C gene expression. The basal transcriptional mechanism was elucidated by Marinovic et al. first and then by the research group led by Bianchi, highlighting the importance of two ubiquitously expressed transcription factors in driving UbC gene expression: Sp1 and YY1 (“Figure 3”).

Figure 3. Ubiquitin C proximal promoter control region with functionally relevant transcription factor binding sites. The region contains 371 nt promoter sequence, the first untranslated exon (63 nt) and the unique intron (812 nt). The transcription start site, TATA-box and intron-exon borders were identified by Nenoi et al. in 1996 [30]. The upstream Sp1 binding sites were characterized by Marinovic et al. [31]. PPARb binding site was identified by Kim [36]. YY1 intronic binding sites were identified by Bianchi et al. [39].
A central role is also given to the unique intron which harbours the two YY1 binding sites. On the other hand, there is a great bulk of non-molecular evidences heightening UbC as one of the most important stress-inducible genes, but a molecular study is still needed to address how this induction is achieved.
6. ACKNOWLEDGEMENTS
We gratefully thank Daniele Tontini for the precious support in figure drawing.
NOTES